Abstract
Background: Ideal dose of the antimicrobials should be decided by considering their killing dynamics since sufficient elimination of the causative microorganisms is critical for proper antimicrobial treatment. In this study, the bactericidal activities of carbapenems by resistance mechanisms were assessed for carbapenem-resistant Pseudomonas aeruginosa. Methods: Minimal inhibition concentrations (MICs) of carbapenems were determined by broth dilution method and the resistance mechanisms were identified by PCR and DNA sequencing. The expression levels of efflux pumps were determined by reverse transcriptase real-time PCR. Time-kill curves were plotted by time-course numeration of the viable cells grown under imipenem and meropenem at 1× and 4× MICs, respectively. Results: One P. aeruginosa strain was susceptible, whereas three were resistant to carbapenems by defective OprD, efflux pump overproduction, and/or IMP-6 production. The susceptible strain had imipenem and meropenem MICs of 2 and 1 mg/L, respectively. The MICs were elevated by eight-fold by defective OprD, 16- and 32-fold by the pump overproduction, and four- and <64-fold by the combination of two determinants and the IMP-6 carbapenemase. While both the carbapenems showed time-dependent bactericidal activity to the susceptible isolate, either of the carbapenem-resistant determinants, such as decreased membrane permeability, carbapenemase production, or the defective OprD,presented concentration-dependent bacteriostatic activity.Conclusion: Different killing dynamics of the carbapenems were observed depending on the resistance determinants, and the results would guide a proper treatment strategy for the patients using these drugs.
Keywords
Bactericidal Carbapenems Killing dynamics Pseudomonas aeruginosa
Figures & Tables
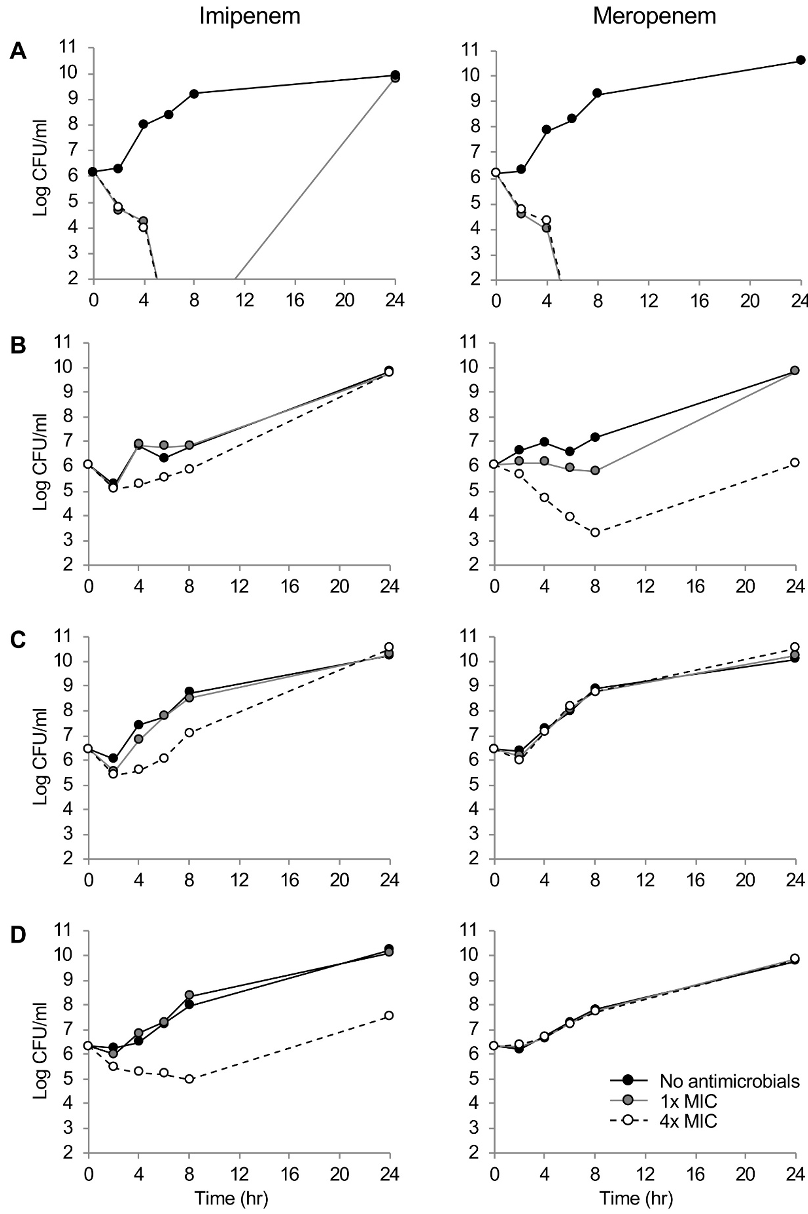
Figure 1. Time–kill curves of imipenem and meropenem against P. aeruginosa (A) A1701DS, (B) B1707∆oprD, (C) F1709 pump↑, (D) B1605∆oprD/ pump↑/IMP strains. Number of bacterial colony forming units are presented to the detection limit (102 CFU/ml). Black circle and black solid line, no antibiotics treated; gray circle and gray solid line, 1×MIC; and open circle and black broken line, 4×MIC.
Figures & Tables
Table 1. Carbapenem susceptibility and relative level of efflux pump gene expression in P. aeroginosa strains
| P. aeruginosa | MIC (mg/L) of* | Relative level of expression to A1701DS† | |||
| Imipenem | Meropenem | MexAB-OprM | MexXY-OprM | MexCD-OprJ | |
| A1701DS | 2 | 1 | 1.0 | 1.0 | 1.0 |
| B1707∆oprD | 16 | 8 | 0.6 | 0.3 | -‡ |
| F1709pump↑ | 32 | 32 | 1.4 | 0.6 | 0.7 |
| B1605∆oprD/ pump↑IMP | 8 | >64 | 1.6 | 3.4 | 2.8 |


