Hui-Jin Yu, Tae Yeul Kim, Eun Jeong Won, Hee Jae Huh
Ann Clin Microbiol 2026 June, 29(2):6. Published on 8 May 2026.
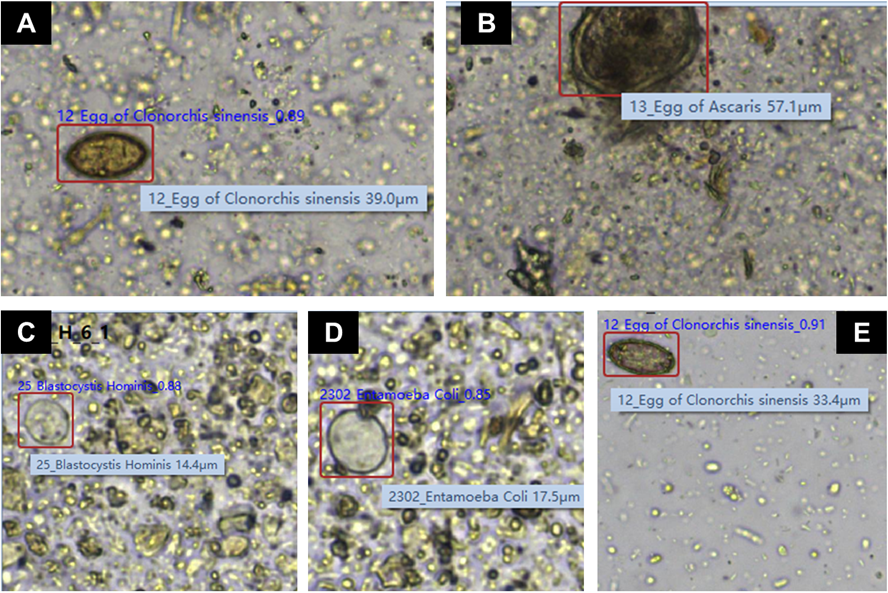
Kibum Jeon, Nuri Lee, Hyun Soo Kim, Han-Sung Kim, Wonkeun Song, Jae-Seok Kim
Ann Clin Microbiol 2026 March, 29(1):1. Published on 6 March 2026.
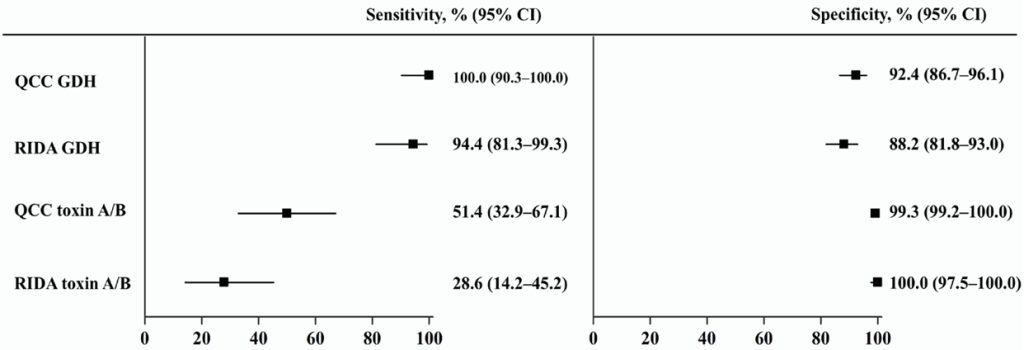
Background: Enzyme immunoassays (EIAs), which detect glutamate dehydrogenase (GDH) and toxin A/B, are widely used to screen for Clostridioides difficile infection (CDI); however, their sensitivity is lower than that of molecular assays. This study aimed to evaluate the performance of two EIAs, C. Diff Quik Chek Complete (QCC) and RIDASCREEN (RIDA), and investigate the cycle threshold (Ct) values from two real-time polymerase chain reaction (PCR) assays (Allplex GI–Bacteria(I) and Xpert C. difficile) in EIA-discordant samples.
Methods: A total of 180 clinical stool samples were tested using QCC, RIDA, and Allplex GI-Bacteria(I) PCR assays. The Xpert C. difficile assay was used to analyze discordant results.
Results: QCC and RIDA showed high sensitivities for GDH detection, 100.0% and 94.4%, respectively. QCC was significantly more sensitive than RIDA for toxin detection (51.4% vs. 28.6%, p = 0.007). In 25 EIA-discordant, Xpert positive samples, the Ct values of the toxin B gene ranged from 31.5 to 44.8 (mean, 38.1) for Allplex PCR and from 23.7 to 36.3 (mean, 30.4) for Xpert PCR. The Ct values of the two PCR assays were not significantly correlated (r = 0.201, p = 0.324).
Conclusion: QCC is a suitable initial immunological test for diagnosing CDI. The lack of correlation in the Ct values between the two real-time PCR assays suggests that assay-specific validation is necessary for cutoff level interpretation.
Kibum Jeon, Nuri Lee, Hyun Soo Kim, Han-Sung Kim, Wonkeun Song, Jae-Seok Kim
Ann Clin Microbiol 2026 March, 29(1):1. Published on 6 March 2026.
Background: Enzyme immunoassays (EIAs), which detect glutamate dehydrogenase (GDH) and toxin A/B, are widely used to screen for Clostridioides difficile infection (CDI); however, their sensitivity is lower than that of molecular assays. This study aimed to evaluate the performance of two EIAs, C. Diff Quik Chek Complete (QCC) and RIDASCREEN (RIDA), and investigate the cycle threshold (Ct) values from two real-time polymerase chain reaction (PCR) assays (Allplex GI–Bacteria(I) and Xpert C. difficile) in EIA-discordant samples.
Methods: A total of 180 clinical stool samples were tested using QCC, RIDA, and Allplex GI-Bacteria(I) PCR assays. The Xpert C. difficile assay was used to analyze discordant results.
Results: QCC and RIDA showed high sensitivities for GDH detection, 100.0% and 94.4%, respectively. QCC was significantly more sensitive than RIDA for toxin detection (51.4% vs. 28.6%, p = 0.007). In 25 EIA-discordant, Xpert positive samples, the Ct values of the toxin B gene ranged from 31.5 to 44.8 (mean, 38.1) for Allplex PCR and from 23.7 to 36.3 (mean, 30.4) for Xpert PCR. The Ct values of the two PCR assays were not significantly correlated (r = 0.201, p = 0.324).
Conclusion: QCC is a suitable initial immunological test for diagnosing CDI. The lack of correlation in the Ct values between the two real-time PCR assays suggests that assay-specific validation is necessary for cutoff level interpretation.
Jungmi Kim, Inyoung Kang, Sunjoo Kim
Ann Clin Microbiol 2026 March, 29(1):3. Published on 9 March 2026.
Background: Environmental contamination with multidrug-resistant organisms (MDROs), including carbapenem-resistant Acinetobacter baumannii (CRAB) and multidrug-resistant Pseudomonas aeruginosa (MRPA), remains a major challenge in healthcare facilities. Hypochlorous acid water (HOCl) has emerged as a promising disinfectant owing to its strong antimicrobial activity and favorable safety profile. This study aimed to evaluate the bactericidal efficacy of atomized HOCl against CRAB and MRPA in a hospital room.
Methods: An atomization experiment was conducted in a two-bed room. CRAB and MRPA were prepared using drying and non-drying methods, respectively. HOCl (CLFine) at concentration of 40 and 300 ppm was atomized using ultrasonic humidifiers. Bacterial samples were collected at 0, 1, 3, and 5 h after atomization. Viable bacterial counts were determined by culture, and bactericidal efficacy was evaluated.
Results: Atomized HOCl exhibited time- and concentration-dependent bactericidal effects against CRAB and MRPA. CRAB and MRPA reached their limits of detection at 3 and 5 h post-atomization at 40 ppm, and at 1 and 3 h at 300 ppm, respectively.
Conclusion: Atomized HOCl effectively inactivated CRAB and MRPA in a hospital room within 3–5 h. These findings support the potential application of HOCl atomization as an adjunctive environmental disinfection strategy for controlling MDRO contamination in healthcare facilities.
Kyeong Seon Ryu, Jae Hyeon Park, Jee-Soo Lee, Moon-Woo Seong
Ann Clin Microbiol 2026 March, 29(1):5. Published on 20 March 2026.
Background: The accurate laboratory-based detection of respiratory viruses, including severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), is essential for effective patient management and infection control in the post–coronavirus disease 2019 era. In this study, the performance of the PowerChek Respiratory Virus Panels 1–4 assay (Kogene Biotech) in detecting non-SARS-CoV-2 respiratory viruses and SARS-CoV-2 was compared with those of two established commercial assays.
Methods: Residual clinical respiratory specimens collected between December 2023 and February 2024 were retested using the PowerChek assay. Of the 129 specimens retested, 80 had tested positive for 14 non-SARS-CoV-2 respiratory virus targets in the Allplex Respiratory Panels 1–3 assay (Seegene) and 49 had been tested using the Alinity m SARS-CoV-2 assay (Abbott; 20 positive and 29 negative for SARS-CoV-2). Agreement, Cohen’s kappa, and discordant results were assessed.
Results: For the 14 non-SARS-CoV-2 respiratory virus targets, the virus-specific overall percent agreement (OPA) ranged from 95.0% to 100%, the positive percent agreement (PPA) ranged from 66.7% to 100%, and the negative percent agreement (NPA) ranged from 96.0% to 100%, with kappa values of 0.64–1.00. Lower agreements were observed for human coronavirus OC43 and influenza A virus. For SARS-CoV-2 detection, the OPA was 98.0% (PPA, 95.0%; NPA, 100%; kappa, 0.96).
Conclusion: The detection performance of the PowerChek assay was comparable to those of the established assays for most respiratory virus targets, with lower agreement observed for some targets. These features support its practical utility for the routine multiplex molecular detection of respiratory viruses, including SARS-CoV-2.
Suyeon Jo, Byoungkuk Kim, Boyeong Kwon, Kwanghyun Kim, Minhyeok Kim, Kihyung Park
Ann Clin Microbiol 2026 March, 29(1):4. Published on 11 March 2026.
Background: Pathogenic Escherichia coli (PEC) is a leading cause of acute diarrhea worldwide. With increasing antibiotic use, antimicrobial resistance in PEC has become a significant public health concern. In this study, we investigated the distribution and antimicrobial susceptibility patterns of PEC isolated from patients with acute diarrhea in the Busan and Gyeongnam regions of South Korea.
Methods: In this retrospective cross-sectional laboratory-based study, 272 PCR-positive residual stool or rectal swab specimens collected between October 2023 and June 2024 were screened; after exclusion of 80 specimens without recoverable or confirmable isolates, 192 pathogenic E. coli isolates were included for pathotype identification and antimicrobial susceptibility testing. Pathotypes were identified using multiplex real-time polymerase chain reaction, and antimicrobial susceptibility was tested against 18 agents using the VITEK II system.
Results: Enteroaggregative E. coli (EAEC) was the most prevalent pathotype (44.8%), followed by enteropathogenic E. coli (34.9%). High resistance rates were observed for ampicillin (77.1%), cefazolin (55.2%), and amoxicillin/clavulanic acid (44.3%). EAEC demonstrated the highest multidrug resistance (MDR) rate (34.9%). Pediatric patients (< 18 years) exhibited significantly higher resistance rates and MDR frequencies compared with adults, particularly against β-lactam antibiotics.
Conclusion: EAEC was identified as the predominant and most resistant pathotype in this region. The elevated resistance levels in children and the prevalence of MDR EAEC underscore the need for continuous local surveillance and appropriate antibiotic stewardship.
Jaesoon Kim, Nakwon Kwak, Jee-Soo Lee, Taek Soo Kim, Moon-Woo Seong, Jae Hyeon Park
Ann Clin Microbiol 2026 March, 29(1):2. Published on 27 February 2026.
For Authors
Publisher
Subscribers
for quarterly newsletter subscription & instant discount on publication fee!
Sponsors