1Department of Laboratory Medicine, Seoul National University Hospital, Seoul, Korea
2Department of Laboratory Medicine, Seoul National University College of Medicine, Seoul, Korea
3Division of Pulmonary and Critical Care Medicine, Department of Internal Medicine, Seoul National University Hospital, Seoul, Korea
4Department of Internal Medicine, Seoul National University College of Medicine, Seoul, Korea
*These authors contributed equally to this work.
Correspondence to Jae Hyeon Park, E-mail: bjack9@snu.ac.kr
Ann Clin Microbiol 2026;29(1):2. https://doi.org/10.5145/ACM.2026.29.1.2
Received on 1 January 2026, Revised on 15 January 2026, Accepted on 30 January 2026, Published on 3 March 2026.
Copyright © Korean Society of Clinical Microbiology.
This is an Open Access article which is freely available under the Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License (CC BY-NC-ND) (https://creativecommons.org/licenses/by-nc-nd/4.0/).
Dear Editor,
Segniliparus spp. are acid-fast, rapidly growing bacteria associated with pulmonary infections, particularly in patients with underlying lung disease. Two species, Segniliparus rugosus and Segniliparus rotundus, were first reported in 2005 from unidentifiable mycobacterial samples suspected of being involved in respiratory mycobacteriosis [1]. Segniliparus spp. infection in humans is exceedingly rare. To date, eight cases have been reported worldwide, with most having been caused by S. rugosus in patients with cystic fibrosis or bronchiectasis [2–7]. In previous reports, species identification relied primarily on 16S rRNA gene sequencing and less frequently on rpoB gene sequencing. Here, we present a case of pulmonary S. rugosus infection in a patient without evidence of systemic immunodeficiency but with bronchiectasis and prior nontuberculous mycobacterial pulmonary disease (NTM-PD), identified for the first time by matrix-assisted laser desorption ionization time-of-flight mass spectrometry (MALDI-TOF MS) and confirmed by whole-genome sequencing (WGS). This case highlights that MALDI-TOF MS can identify rare acid-fast organisms within routine clinical workflows, with WGS providing species-level confirmation.
A 66-year-old woman presented with worsening cough and purulent sputum for 6 months. She had no evidence of systemic immunodeficiency (HIV-negative, no history of malignancy or immunosuppressive therapy). She had been diagnosed with NTM-PD caused by Mycobacterium avium 2 years earlier. For suspected M. avium complex pulmonary disease, empirical treatment comprising clarithromycin, ethambutol, and rifampin was initiated at another institution following positive acid-fast bacillus (AFB) staining. When the patient failed to respond to the treatment after 10 weeks and was referred to our institution, a new regimen consisting of azithromycin, ethambutol, and clofazimine was initiated. After 5 months without clinical improvement, all antibiotics were discontinued because she developed azotemia and IgA nephropathy.
After antibiotic discontinuation, serial sputum samples were strongly AFB-positive, and occasional growth was observed in BD BACTEC MGIT broth (Becton, Dickinson and Company) and on Ogawa agar (Fig. 1A). Isolates from these cultures were also strongly AFB-positive (Fig. 1B). However, because repeated Allplex TB/NTM real-time PCR results (Seegene) were negative, the isolates were initially regarded as contaminants. On a colony smear, the organism showed strongly acid-fast, short, nonbranching rods.
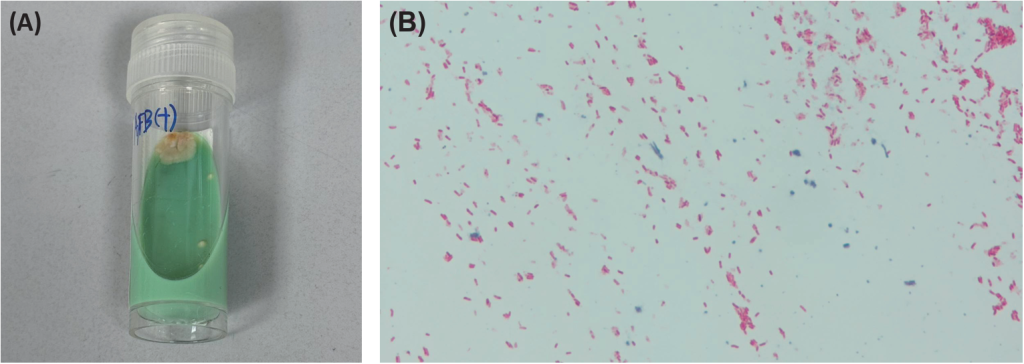
Fig. 1. Colony morphology and acid-fast staining of Segniliparus rugosus. (A) Colonies on Ogawa medium. Rough, dry, buff-to-orange colonies. (B) Acid-fast bacillus stain (colony smear). Strongly acid-fast short rods were observed, unlike Nocardia spp., which are typically weakly/partially acid-fast and show branching filaments.
When the patient’s renal function improved but respiratory symptoms and CT findings worsened, antibiotic therapy consisting of azithromycin, ethambutol, and clofazimine was resumed after a 10-month hiatus. After 3 months of minimal symptomatic improvement, colonies previously presumed to be contaminants were subcultured on blood agar and analyzed using MALDI-TOF MS to identify the causative organism. The colonies were processed using a direct transfer method and identified as S. rugosus (score: 2.09) using a Bruker Biotyper and MALDI Biotyper Compass Library (DB12348, version 12.0; Bruker Daltonics).
The patient was admitted, and empirical antibiotic therapy—aligned with regimens for rapidly growing mycobacteria—was initiated, comprising azithromycin, clofazimine, amikacin, and imipenem. The symptoms improved dramatically within a few days of therapy initiation, and management was subsequently continued on an outpatient basis. Drug susceptibility testing (DST) was initially attempted with the Sensititre RAPMYCOI plate (Thermo Fisher Scientific) in cation-adjusted Mueller–Hinton broth, but the isolate failed to grow. In line with prior reports on Segniliparus spp., DST was repeated in Middlebrook 7H9 broth (Korean Institute of Tuberculosis, Cheongju, Korea; Table 1). Amikacin was replaced with ciprofloxacin 3 months later, in light of the DST results and the patient’s renal function.
Table 1. Antimicrobial susceptibility of the S. rugosus clinical isolate
| Antimicrobial agent | MIC (μg/mL) | Interpretation a) |
|---|---|---|
| Clarithromycin | > 32 | R |
| Amikacin | > 128 | R |
| Moxifloxacin | 2 | I |
| Linezolid | 16 | I |
| Streptomycin | 64 | N/A |
| Ciprofloxacin | ≤ 0.25 | S |
| Doxycycline | > 16 | R |
| Clofazimine | ≤ 0.25 | N/A |
| Trimethoprim/Sulfamethoxazole | ≤ 0.5/9.5 | S |
| Cefoxitin | 128 | R |
| Imipenem | 4 | S |
| Meropenem | 8 | I |
| Tobramycin | > 8 | R |
| Tigecycline | 4 | N/A |
a)Interpretations were made according to CLSI M62 for rapidly growing mycobacteria and are in line with those reported in previous case studies.
Abbreviations: MIC, minimum inhibitory concentration; R, resistant; I, intermediate; S, susceptible; N/A, no available breakpoint.
WGS was performed using the NovaSeq 6000Dx platform (Illumina). Raw Illumina paired-end reads were adapter- and quality-trimmed with fastp and assembled using SPAdes (isolate mode), and contigs < 200 bp were excluded prior to calculating assembly metrics. Contigs flagged as contaminants by the National Center for Biotechnology Information (NCBI) contamination screen (primate/fungal) were removed prior to public deposition, and all assembly metrics and downstream analyses reported herein were calculated for the decontaminated assembly. The assembled genome measured 3,872,893 bp across 153 contigs (N50: 217,954 bp; largest contig: 410,624 bp; GC content: 68.16%). BUSCO v6.0.0 using the Actinomycetes odb12 lineage (n = 355) showed 95.8% completeness (340/355 complete single-copy), indicating a high-quality assembly. The average nucleotide identity (ANI) was calculated using FastANI v1.34 against the NCBI RefSeq reference genomes of representative Segniliparus strains, yielding 99.61% for S. rugosus (GCF_000185725.2; 1144/1265 fragments mapped, 90.4% coverage) and 81.69% for S. rotundus (GCF_000092825.1; 719/1265 fragments mapped, 56.8% coverage). As these values were above and well below the ~95%–96% species threshold [8], respectively, the isolate was assigned to S. rugosus, consistent with the MALDI-TOF MS results. Beyond the ANI calculated from the assembled genome, phylogenomic analysis using the Type Strain Genome Server (accessed September 5, 2025), based on Genome BLAST Distance Phylogeny, placed the isolate within the S. rugosus clade, clustered with the type strain, and independently corroborated species assignment (Fig. 2).
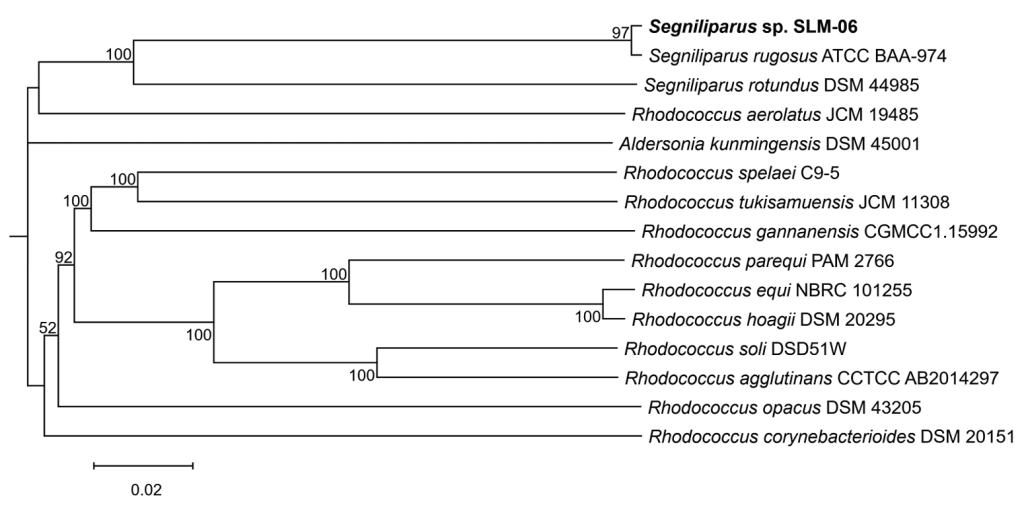

Fig. 2. Phylogenomic placement of the clinical Segniliparus rugosus isolate by Type Strain Genome Server (TYGS). A genome-based tree was generated with the TYGS (accessed September 5, 2025) using Genome BLAST Distance Phylogeny (GBDP) distances and FastME. The isolate clustered within the S. rugosus clade together with the type strain, supporting the species assignment. The tree was visualized using iTOL v7. Node labels indicate GBDP pseudo-bootstrap support values (only ≥ 50% are shown), and the scale bar indicates GBDP distance.
Here, we describe a case of pulmonary disease caused by S. rugosus infection in a patient without evidence of systemic immunodeficiency but with bronchiectasis and prior NTM-PD. All previous reports of Segniliparus infections involved patients with either cystic fibrosis or bronchiectasis, both of which predispose patients to nontuberculous mycobacterial disease [9]. Although further investigation is required, we believe that such structural lung damage and impaired mucociliary clearance can predispose patients to infection by rare pathogens, such as Segniliparus spp. As our patient had a history of NTM-PD and radiologically identifiable bronchiectasis, she may have shared these risk factors.
As summarized in Table 2, most reported S. rugosus infections occurred in patients with cystic fibrosis or bronchiectasis, and the diagnosis relied primarily on 16S rRNA or, in one case, rpoB sequencing. To the best of our knowledge, no previous case has identified S. rugosus by MALDI-TOF MS or confirmed its identification by WGS, expanding the spectrum of susceptible hosts to patients with prior NTM-PD. Despite limited reference spectra in the Bruker MALDI Biotyper Compass library (DB12348, version 12.0), the isolate was identified as S. rugosus with a high-confidence score (2.09; ≥ 2.0 generally supports species-level identification).
Table 2. Human infections due to Segniliparus spp., including the present case, with identification method, risk factors, DST, and outcomes
| Author (year) | Species | Identification method | Risk factor | DST | Outcome |
|---|---|---|---|---|---|
| Butler et al. (2007) [2] | S. rugosus (3 cases) | 16S rRNA sequencing | CF | BMD in Middlebrook 7H9 broth | Treated |
| Hansen et al. (2009) [3] | S. rugosus | 16S rRNA sequencing | CF | Sensititre RAPMYCOI in Middlebrook 7H9 broth | Treated |
| Koh et al. (2011) [4] | S. rotundus | 16S rRNA and rpoB sequencing | Non-CF bronchiectasis | Done | Treated |
| Choi et al. (2012) [5] | S. rugosus | 16S rRNA sequencing | Non-CF bronchiectasis | Done | Not treated |
| Lee et al. (2014) [6] | S. rugosus | 16S rRNA sequencing | Non-CF bronchiectasis | Failed | Treated |
| Zurita et al. (2020) [7] | S. rugosus | 16S rRNA sequencing | CF | Sensititre RAPMYCOI | Not reported |
| Present case | S. rugosus | MALDI-TOF MS and WGS | Previous NTM-PD | BMD in Middlebrook 7H9 broth | Treated |
Abbreviations: DST, drug susceptibility testing; CF, cystic fibrosis; BMD, broth microdilution; MALDI-TOF MS, matrix-assisted laser desorption ionization time-of-flight mass spectrometry; NTM-PD, nontuberculous mycobacterial pulmonary disease; WGS, whole-genome sequencing.
AFB-positive but TB/NTM PCR-negative isolates are a recurring diagnostic pitfall and may be dismissed as contaminants. Therefore, we suggest a pragmatic workflow: retain AFB-positive, PCR-negative cultures, perform routine subculture, and apply MALDI-TOF MS to any growth before discarding it as contamination. WGS-based confirmation (e.g., ANI and phylogenomics) can be reserved for selected cases with therapeutic implications or clinic–laboratory discordance. This blind spot is not unique to Segniliparus, as other nonmycobacterial acid-fast taxa can also yield culture-positive, PCR-negative results, underscoring the need for culture-based identification along with molecular testing [10].
In terms of antimicrobial therapy, previous reports described variable responses to imipenem, rifabutin, moxifloxacin, and trimethoprim-sulfamethoxazole [2,3]. More recent cases have adopted regimens aligned with treatment guidelines for rapidly growing mycobacteria, typically combining a macrolide with a f luoroquinolone and a parenteral agent, such as amikacin or imipenem [6]. In the present case, dramatic clinical improvement was achieved with azithromycin, clofazimine, amikacin, and imipenem, although amikacin was later replaced with ciprofloxacin owing to high minimum inhibitory concentrations and renal dysfunction. As reflected in Table 2, treatment outcomes in earlier cases have been heterogeneous, likely owing to the rarity of the organism and the absence of standardized regimens. Our findings emphasize the importance of performing DST under appropriate culture conditions and tailoring therapy according to both microbiological results and patient comorbidities. Such individualized management is particularly relevant for rare organisms, such as S. rugosus, for which evidence-based treatment guidance remains limited.
This case highlights that MALDI-TOF MS can rapidly identify S. rugosus from AFB-positive TB/NTM PCR-negative isolates that might otherwise be dismissed as contaminants, while WGS provides species-level confirmation when clinically indicated. Adopting this stepwise approach may shorten identification time and support individualized therapies for rare acid-fast infections.
Written informed consent was obtained from the patient for the publication of this case report and the accompanying images.
No potential conflicts of interest relevant to this article were reported.
None.
The whole-genome shotgun (WGS) project has been deposited at GenBank under the WGS master accession JBQWBN000000000.1. Raw sequencing reads are available in the NCBI Sequence Read Archive (SRA) under BioProject PRJNA1313124 (BioSample SAMN50884781).
1. Butler WR, Floyd MM, Brown JM, Toney SR, Daneshvar MI, Cooksey RC, et al. Novel mycolic acid-containing bacteria in the family Segniliparaceae fam. nov., including the genus Segniliparus gen. nov., with descriptions of Segniliparus rotundus sp. nov. and Segniliparus rugosus sp. nov. Int J Syst Evol Microbiol 2005;55:1615-24.

2. Butler WR, Sheils CA, Brown-Elliott BA, Charles N, Colin AA, Gant MJ, et al. First isolations of Segniliparus rugosus from patients with cystic fibrosis. J Clin Microbiol 2007;45:3449-52.


3. Hansen T, Van-Kerckhof J, Jelfs P, Wainwright C, Ryan P, Coulter C. Segniliparus rugosus infection, Australia. Emerg Infect Dis 2009;15:611-3.


4. Koh WJ, Choi GE, Lee SH, Park YK, Lee NY, Shin SJ. First case of Segniliparus rotundus pneumonia in a patient with bronchiectasis. J Clin Microbiol 2011;49:3403-5.


5. Choi SM, Kang HJ, Jeong YJ, Lim JH, Choe WS, Hwang SH, et al. First isolation of Segniliparus rugosus from a patient with radiologic features similar to non-tuberculous mycobacteriosis. Tuberc Respir Dis (Seoul) 2012;72:82-7.
6. Lee JY, Chon GR, Jung TY, Sung H, Shim TS, Jo KW. A case of Segniliparus rugosus pulmonary infection in an immunocompetent patient with non-cystic fibrosis. Tuberc Respir Dis (Seoul) 2014;77:227-9.

7. Zurita J, Sevillano G, González C, Lascano Y. Segniliparus rugosus from the sputum of a child with cystic fibrosis in Ecuador: challenges in bacterial identification. New Microbes New Infect 2020;35:100668.


8. Riesco R and Trujillo ME. Update on the proposed minimal standards for the use of genome data for the taxonomy of prokaryotes. Int J Syst Evol Microbiol 2024;74:006300.


9. Van Braeckel E and Bosteels C. Growing from common ground: nontuberculous mycobacteria and bronchiectasis. Eur Respir Rev 2024;33:240058.


10. Mochon AB, Sussland D, Saubolle MA. Aerobic Actinomycetes of clinical significance. Microbiol Spectr 2016;4:10.1128/microbiolspec.DMIH2-0021-2015.

1. Butler WR, Floyd MM, Brown JM, Toney SR, Daneshvar MI, Cooksey RC, et al. Novel mycolic acid-containing bacteria in the family Segniliparaceae fam. nov., including the genus Segniliparus gen. nov., with descriptions of Segniliparus rotundus sp. nov. and Segniliparus rugosus sp. nov. Int J Syst Evol Microbiol 2005;55:1615-24.

2. Butler WR, Sheils CA, Brown-Elliott BA, Charles N, Colin AA, Gant MJ, et al. First isolations of Segniliparus rugosus from patients with cystic fibrosis. J Clin Microbiol 2007;45:3449-52.


3. Hansen T, Van-Kerckhof J, Jelfs P, Wainwright C, Ryan P, Coulter C. Segniliparus rugosus infection, Australia. Emerg Infect Dis 2009;15:611-3.


4. Koh WJ, Choi GE, Lee SH, Park YK, Lee NY, Shin SJ. First case of Segniliparus rotundus pneumonia in a patient with bronchiectasis. J Clin Microbiol 2011;49:3403-5.


5. Choi SM, Kang HJ, Jeong YJ, Lim JH, Choe WS, Hwang SH, et al. First isolation of Segniliparus rugosus from a patient with radiologic features similar to non-tuberculous mycobacteriosis. Tuberc Respir Dis (Seoul) 2012;72:82-7.
6. Lee JY, Chon GR, Jung TY, Sung H, Shim TS, Jo KW. A case of Segniliparus rugosus pulmonary infection in an immunocompetent patient with non-cystic fibrosis. Tuberc Respir Dis (Seoul) 2014;77:227-9.

7. Zurita J, Sevillano G, González C, Lascano Y. Segniliparus rugosus from the sputum of a child with cystic fibrosis in Ecuador: challenges in bacterial identification. New Microbes New Infect 2020;35:100668.


8. Riesco R and Trujillo ME. Update on the proposed minimal standards for the use of genome data for the taxonomy of prokaryotes. Int J Syst Evol Microbiol 2024;74:006300.


9. Van Braeckel E and Bosteels C. Growing from common ground: nontuberculous mycobacteria and bronchiectasis. Eur Respir Rev 2024;33:240058.


10. Mochon AB, Sussland D, Saubolle MA. Aerobic Actinomycetes of clinical significance. Microbiol Spectr 2016;4:10.1128/microbiolspec.DMIH2-0021-2015.
