Department of Laboratory Medicine, Seegene Medical Foundation, Busan, Korea
Correspondence to Suyeon Jo E-mail: freedom96@naver.com; Kihyung Park E-mail: p0413@mf.seegene.com
Ann Clin Microbiol 2026;29(1):4. https://doi.org/10.5145/ACM.2026.29.1.4
Received on 12 January 2026, Revised on 26 February 2026, Accepted on 27 February 2026, Published on 11 March 2026.
Copyright © Korean Society of Clinical Microbiology.
This is an Open Access article which is freely available under the Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License (CC BY-NC-ND) (https://creativecommons.org/licenses/by-nc-nd/4.0/).
Background: Pathogenic Escherichia coli (PEC) is a leading cause of acute diarrhea worldwide. With increasing antibiotic use, antimicrobial resistance in PEC has become a significant public health concern. In this study, we investigated the distribution and antimicrobial susceptibility patterns of PEC isolated from patients with acute diarrhea in the Busan and Gyeongnam regions of South Korea.
Methods: In this retrospective cross-sectional laboratory-based study, 272 PCR-positive residual stool or rectal swab specimens collected between October 2023 and June 2024 were screened; after exclusion of 80 specimens without recoverable or confirmable isolates, 192 pathogenic E. coli isolates were included for pathotype identification and antimicrobial susceptibility testing. Pathotypes were identified using multiplex real-time polymerase chain reaction, and antimicrobial susceptibility was tested against 18 agents using the VITEK II system.
Results: Enteroaggregative E. coli (EAEC) was the most prevalent pathotype (44.8%), followed by enteropathogenic E. coli (34.9%). High resistance rates were observed for ampicillin (77.1%), cefazolin (55.2%), and amoxicillin/clavulanic acid (44.3%). EAEC demonstrated the highest multidrug resistance (MDR) rate (34.9%). Pediatric patients (< 18 years) exhibited significantly higher resistance rates and MDR frequencies compared with adults, particularly against β-lactam antibiotics.
Conclusion: EAEC was identified as the predominant and most resistant pathotype in this region. The elevated resistance levels in children and the prevalence of MDR EAEC underscore the need for continuous local surveillance and appropriate antibiotic stewardship.
Antimicrobial stewardship, Bacterial drug resistance, Diarrhea, Escherichia coli, Republic of Korea
Acute diarrhea is a globally important public health issue and a major cause of death among children aged ≤ 5 years. Approximately 443,800 children die from acute diarrhea annually, with over 1.7 billion cases reported worldwide among children each year [1]. Acute diarrhea can last for several days, causing dehydration and electrolyte loss and, in severe cases, leading to sepsis. Although the mortality rate among adults is relatively low, the socioeconomic burden can be substantial due to increased healthcare costs and progression to chronic bowel disease. Bacterial diarrhea may impair immune function and carries a high risk of complications in elderly individuals [2]. Recently, the incidence of bacterial enterocolitis among causative pathogens of acute diarrhea has increased. According to reports of grade 4 bacterial enterocolitis from 206 sentinel surveillance centers in South Korea in 2023, salmonellosis was the most common (3,540 cases), followed by campylobacteriosis (3,167 cases), enteropathogenic Escherichia coli (EPEC) (1,963 cases), and enterotoxigenic E. coli (ETEC) (481 cases). Pathogenic E. coli (PEC) infections increased rapidly, showing a 2.5-fold rise from 1,068 cases in 2022 to 2,499 cases in 2023. In addition, during the 2024–2025 period, campylobacteriosis cases progressively decreased, whereas salmonellosis and PEC infections continued to increase [3]. These findings indicate that PEC is firmly established as a major cause of acute diarrhea in South Korea.
Although PEC infections usually resolve spontaneously, antibiotic treatment may be required for high-risk groups or for individuals infected with certain pathotypes. The rising rate of antibiotic resistance among PEC is closely associated with increased antibiotic use. Specifically, intensive antibiotic use in healthcare settings fosters an environment in which isolated E. coli strains can more readily develop or acquire resistance. These resistant strains may subsequently disseminate into the community, increasing the carriage rate of multidrug-resistant E. coli among the general population [4]. According to reports from South Korea, PEC shows high resistance rates to major antibiotics, including nalidixic acid (44.0%), tetracycline (41.3%), ampicillin (AMP, 40.0%), and sulfamethoxazole/trimethoprim (SXT, 34.7%) [5,6]. These results show that, in addition to the increasing incidence of PEC infections, antibiotic resistance is emerging as a clinically important problem. This trend is likely to restrict antibiotic treatment options and increase the burden of infection control. Hence, analyzing antimicrobial susceptibility patterns among community-derived PEC isolates is important for generating baseline data to monitor the spread of resistance and to support guidelines for rational antibiotic use and treatment.
In this study, we aimed to analyze the antibiotic resistance profiles of PEC isolates from patients with acute diarrhea in southeastern South Korea. The primary objective was to describe antimicrobial resistance profiles of PEC isolates. Secondary objectives were to compare resistance by pathotype and age group and to describe annual changes in pathotype distribution. The findings of this study are intended to provide baseline data for monitoring regional antibiotic resistance trends and to inform appropriate empirical antimicrobial therapy.
This was a retrospective cross-sectional laboratory-based observational study of pathogenic Escherichia coli isolates from patients with acute diarrhea in Busan and Gyeongsangnam-do, Korea.
This study included 272 PEC-positive stool and rectal swab samples collected from patients at hospitals and public health centers in Busan and Gyeongsangnam-do between October 2023 and June 2024.
Eligible specimens were residual stool or rectal swab samples submitted for diagnostic evaluation of acute diarrhea. When multiple specimens were available from the same patient, only the first eligible isolate was included. Of these, 80 samples were excluded: 64 were initially identified as E. coli-positive during Multiplex real-time polymerase chain reaction (PCR) using the Allplex GI-Bacteria II Assay (Seegene) for the pathogen causing of acute diarrhea but showed no growth on subculture for antimicrobial susceptibility testing, and 16 showed no amplification of E. coli genes on confirmatory testing using subcultured colonies. Thus, 192 samples were included in the final analysis.
The primary outcome variables were antibiotic resistance to each tested antimicrobial agent and multidrug resistance (MDR) among pathogenic Escherichia coli isolates. Antibiotic susceptibility results were categorized as resistant, intermediate, susceptible, or susceptible-dose-dependent according to Clinical and Laboratory Standards Institute criteria; for comparative analyses, resistance rates were calculated as the proportion of isolates classified as resistant for each agent. MDR was defined as resistance to 3 or more antimicrobial classes. The main explanatory variables were E. coli pathotype, classified as enteroaggregative E. coli (EAEC), enteropathogenic E. coli (EPEC), enterotoxigenic E. coli (ETEC), Shiga toxin-producing E. coli (STEC), E. coli O157, and EAEC/EPEC double positivity, and age group, categorized as pediatric (<18 years) or adult (≥18 years). Time period was additionally analyzed as a categorical variable to compare annual changes in pathotype distribution. Samples simultaneously positive for STEC, E. coli O157, and EAEC/EPEC were excluded from MDR subgroup analyses because of the small sample size.
Residual samples were stored at 2–8°C (up to 24 h) and inoculated onto MacConkey agar (BD Difco) using a cotton swab.
After incubation for 24 h at 35°C in a 5% CO2 atmosphere, lactose-fermenting colonies appearing pink on MacConkey agar were considered presumptive E. coli. Representative colonies showing typical colony morphology were selected and subcultured onto blood agar (BD Difco) and incubated under the same conditions for an additional 24 h. When multiple colony morphotypes were observed, one representative colony from each morphotype was selected for further analysis.
For confirmatory testing, subcultured colonies were suspended in 0.45% physiological saline adjusted to a 0.5 McFarland standard (approximately 3 mL). Multiplex real-time PCR was performed to identify specific pathotypes. This assay targets the eaeA gene of EPEC, lt and st genes of ETEC, the aggR gene of enteroaggregative E. coli (EAEC), stx1 and stx2 genes of Shiga toxin-producing E. coli (STEC), and the rfbE gene of E. coli O157.
Nucleic acid extraction was performed using an automated device (Maelstrom 9600; TANBead) with STARMag M96 reagent (Seegene) according to the manufacturer’s guidelines. Amplification was performed using a CFX96 real-time PCR system (Bio-Rad Laboratories). PCR conditions consisted of an initial denaturation at 95°C for 15 min, followed by 46 cycles of denaturation (95°C for 10 s), annealing (60°C for 15 s), and extension (72°C for 15 s). Fluorescence signals were automatically analyzed using Seegene Viewer Software.
E. coli-confirmed isolates were subjected to antimicrobial susceptibility testing using the VITEK II System (bioMérieux) with the AST-N415 card. The card contained the following 18 antibiotics: AMP, amoxicillin/clavulanic acid, piperacillin/tazobactam, cefazolin, cefotaxime, ceftazidime, cefoxitin, cefepime, imipenem, meropenem, ertapenem, gentamicin, amikacin, aztreonam, ciprofloxacin (CIP), SXT, nitrofurantoin, and tigecycline. Isolates freshly subcultured from blood agar were suspended in 0.45% physiological saline, adjusted to a turbidity equivalent to a 0.5 McFarland standard (approximately 3 mL), and analyzed within 1 h. Test results were recorded as resistant, susceptible, intermediate, or susceptible-dose-dependent. The minimum inhibitory concentration of each antibiotic was interpreted according to the Clinical and Laboratory Standards Institute M100 guidelines [7].
Because only PCR-positive specimens yielding recoverable isolates on subculture were included, selection bias toward viable organisms or higher bacterial loads is possible.
The study size was determined by all eligible specimens collected during the prespecified study period; no formal sample-size calculation was performed because this was a retrospective descriptive study.
Descriptive statistics for PEC detection frequency and antibiotic resistance rates were analyzed using Microsoft Excel (Microsoft Corp.). Chi-square tests were used to compare resistance rates to individual antibiotics among different pathotypes (EAEC, EPEC, and ETEC), the prevalence of antibiotic resistance and multidrug resistance (MDR) between pediatric and adult groups, and the annual distribution of PEC pathotypes over the three-year study period. Statistical analyses were performed using MedCalc software (version 22.009; MedCalc Software Ltd.), and P-values < 0.05 were considered statistically significant. An antibiotic resistance rate > 20% was defined as “high resistance,” whereas susceptibility rate ≥ 80% was defined as “high susceptibility” [8]. MDR was defined as resistance to ≥ 3 classes of antibiotics. Samples simultaneously positive for STEC, E. coli O157, and EAEC/EPEC were excluded from the analysis because of the small sample size.
Of the 192 samples confirmed to be PEC-positive, EAEC was the most common pathotype (86 samples, 44.8%), followed by EPEC (67 samples, 34.9%), ETEC (31 samples, 16.1%), STEC (4 samples, 2.1%), E. coli O157 (2 samples, 1.0%), and EAEC/EPEC double positivity (2 samples, 1.0%). The antibiotics with the highest resistance rates, in descending order, were AMP (77.1%), followed by cefazolin (CFZ) (55.2%), amoxicillin/clavulanic acid (AMC) (44.3%), cefotaxime (CTX) (38.0%), SXT (34.9%), and ceftazidime (CAZ) (21.4%; Table 1).
Table 1. Antibiotic resistance profiles of pathogenic Escherichia coli isolates (N = 192)
| Antibiotics | R (n) | S (n) | I (n) | SDD (n) | R (%) | S (%) |
|---|---|---|---|---|---|---|
| AMP | 148 | 42 | 2 | 0 | 77.1 | 22.9 |
| AMC | 85 | 102 | 5 | 0 | 44.3 | 55.7 |
| PT | 30 | 157 | 0 | 5 | 15.6 | 81.8 |
| CFZ | 106 | 86 | 0 | 0 | 55.2 | 44.8 |
| CTX | 73 | 119 | 0 | 0 | 38.0 | 62.0 |
| CAZ | 41 | 113 | 38 | 0 | 21.4 | 58.6 |
| FEP | 11 | 172 | 0 | 9 | 5.7 | 89.6 |
| IPM | 1 | 191 | 0 | 0 | 0.5 | 99.5 |
| MEM | 0 | 192 | 0 | 0 | 0.0 | 100.0 |
| ETP | 0 | 191 | 1 | 0 | 0.0 | 99.5 |
| AMK | 1 | 191 | 0 | 0 | 0.5 | 99.5 |
| GEN | 21 | 170 | 1 | 0 | 10.9 | 89.1 |
| CIP | 29 | 68 | 95 | 0 | 15.1 | 49.5 |
| SXT | 67 | 125 | 0 | 0 | 34.9 | 65.1 |
| NIT | 1 | 190 | 1 | 0 | 0.5 | 99.0 |
| ATM | 4 | 188 | 0 | 0 | 2.1 | 97.9 |
| FOX | 4 | 188 | 0 | 0 | 2.1 | 97.9 |
| TGC | 4 | 188 | 0 | 0 | 2.1 | 97.9 |
Abbreviations: R, resistant; S, susceptible; I, intermediate; SDD, susceptible-dose dependent; AMP, ampicillin; AMC, amoxicillin-clavulanic acid; PT, piperacillin-tazobactam; CFZ, cefazolin; CTX, cefotaxime; CAZ, ceftazidime; FEP, cefepime; IPM, imipenem; MEM, meropenem; ETP, ertapenem; AMK, amikacin; GEN, gentamicin; CIP, ciprofloxacin; SXT, sulfamethoxazole-trimethoprim; NIT, nitrofurantoin; ATM, aztreonam; FOX, cefoxitin; TGC, tigecycline.
Antibiotic resistance rates varied significantly by PEC pathotype, with EAEC exhibiting the highest overall resistance compared to other pathotypes (P < 0.05). Specifically, EAEC showed markedly higher resistance rates than EPEC and ETEC for several key antibiotics, including AMP (94.2% vs. 59.7% and 64.5%), AMC (57.0% vs. 31.3% and 32.2%), and CFZ (69.8% vs. 37.3% and 48.4%, respectively). Furthermore, resistance to CTX and SXT was notably higher in EAEC (51.2% for both) than in the other pathotypes. MDR was identified in 55 of the 192 total isolates (28.6%) and was most prevalent among EAEC isolates (34.9%), followed by EPEC (25.4%) and ETEC (12.9%; P < 0.05) (Table 2).
Table 2. Comparison of antibiotic resistance and multidrug resistance rates among pathogenic Escherichia coli pathotypes
| Variables | EAEC (n = 86) | EPEC (n = 67) | ETEC (n = 31) |
|---|---|---|---|
| Antibiotic resistance rates, n (%) | |||
| AMP | 81 (94.2) | 40 (59.7) | 20 (64.5) |
| AMC | 49 (57.0) | 21 (31.3) | 10 (32.2) |
| PT | 17 (19.8) | 9 (13.4) | 1 (3.2) |
| CFZ | 60 (69.8) | 25 (37.3) | 15 (48.4) |
| CTX | 44 (51.2) | 17 (25.4) | 7 (22.6) |
| CAZ | 25 (29.1) | 9 (13.4) | 3 (9.7) |
| FEP | 5 (5.8) | 2 (3.0) | 1 (3.2) |
| IPM | 0 (0.0) | 1 (1.5) | 0 (0.0) |
| MEM | 0 (0.0) | 0 (0.0) | 0 (0.0) |
| ETP | 0 (0.0) | 0 (0.0) | 0 (0.0) |
| AMK | 0 (0.0) | 1 (1.5) | 0 (0.0) |
| GEN | 4 (4.7) | 11 (16.4) | 3 (9.7) |
| CIP | 9 (10.5) | 12 (17.9) | 5 (16.1) |
| SXT | 44 (51.2) | 15 (22.4) | 6 (19.4) |
| NIT | 0 (0.0) | 0 (0.0) | 0 (0.0) |
| ATM | 2 (2.3) | 1 (1.5) | 0 (0.0) |
| FOX | 2 (2.3) | 1 (1.5) | 0 (0.0) |
| TGC | 2 (2.3) | 1 (1.5) | 0 (0.0) |
| MDR, n (%) | 30 (34.9) | 17 (25.4) | 4 (12.9) |
Abbreviations: EAEC, enteroaggregative Escherichia coli; EPEC, enteropathogenic Escherichia coli; ETEC, enterotoxigenic Escherichia coli; AMP, ampicillin; AMC, amoxicillin-clavulanic acid; PT, piperacillin-tazobactam; CFZ, cefazolin; CTX, cefotaxime; CAZ, ceftazidime; FEP, cefepime; IPM, imipenem; MEM, meropenem; ETP, ertapenem; AMK, amikacin; GEN, gentamicin; CIP, ciprofloxacin; SXT, sulfamethoxazole- trimethoprim; NIT, nitrofurantoin; ATM, aztreonam; FOX, cefoxitin; TGC, tigecycline; MDR, multidrug resistance.
Of the 192 PEC-positive samples, 120 (62.5%) were obtained from children (< 18 years) and 72 (37.5%) from adults. Among pediatric samples, the antibiotics with the highest resistance rates, in descending order, were AMP (81.0%), CFZ (63.6%), AMC (56.2%), CTX (48.8%), SXT (34.7%), CAZ (28.9%), and piperacillin/tazobactam (PT) (23.1%). Among adult samples, the highest resistance rates were observed for AMP (70.4%), CFZ (40.9%), SXT (35.2%), and AMC (23.9%). The MDR rates for EAEC, EPEC, and ETEC isolates in pediatric patients were 43.1%, 31.9%, and 11.1%, respectively, whereas the corresponding rates in adults were 18.5%, 15.0%, and 13.6%, respectively. EAEC and EPEC isolates from children exhibited significantly higher MDR frequencies compared with those from adults (P < 0.05; Table 3).
Table 3. Comparison of antibiotic resistance and multidrug resistance rates of pathogenic Escherichia coli isolates between pediatric and adult patients
| Variables | Pediatrics (n = 120) | Adults (n = 72) |
|---|---|---|
| Antibiotic resistance rates, n (%) | ||
| AMP | 97 (81.0) | 51 (70.4) |
| AMC | 67 (56.2) | 17 (23.9) |
| PT | 28 (23.1) | 2 (2.8) |
| CFZ | 76 (63.6) | 29 (40.9) |
| CTX | 59 (48.8) | 14 (19.4) |
| CAZ | 35 (28.9) | 6 (8.3) |
| FEP | 8 (6.6) | 3 (4.2) |
| IPM | 0 (0.0) | 1 (1.4) |
| MEM | 0 (0.0) | 0 (0.0) |
| ETP | 0 (0.0) | 0 (0.0) |
| AMK | 1 (0.8) | 0 (0.0) |
| GEN | 14 (11.6) | 7 (9.9) |
| CIP | 19 (15.7) | 10 (14.1) |
| SXT | 42 (34.7) | 25 (35.2) |
| NIT | 1 (0.8) | 0 (0.0) |
| ATM | 4 (3.3) | 0 (0.0) |
| FOX | 4 (3.3) | 0 (0.0) |
| TGC | 4 (3.3) | 0 (0.0) |
| MDRa), n (%) | ||
| EAEC | 25 (43.1) | 5 (18.5) |
| EPEC | 15 (31.9) | 3 (15.0) |
| ETEC | 1 (11.1) | 3 (13.6) |
a) Samples simultaneously positive for Shiga toxin-producing E. coli, E. coli O157, and EAEC/EPEC were excluded from the MDR analysis because of the small sample size.
Abbreviations: AMP, ampicillin; AMC, amoxicillin-clavulanic acid; PT, piperacillin-tazobactam; CFZ, cefazolin; CTX, cefotaxime; CAZ, ceftazidime; FEP, cefepime; IPM, imipenem; MEM, meropenem; ETP, ertapenem; AMK, amikacin; GEN, gentamicin; CIP, ciprofloxacin; SXT, sulfamethoxazole- trimethoprim; NIT, nitrofurantoin; ATM, aztreonam; FOX, cefoxitin; TGC, tigecycline; MDR, multidrug resistance; EAEC, enteroaggregative Escherichia coli; EPEC, enteropathogenic Escherichia coli; ETEC, enterotoxigenic Escherichia coli.
The distribution of PEC detected during the study period (October 2023 to June 2024) was 40.0% EAEC, 41.8% EPEC, and 13.6% ETEC. During the same period in the previous year (2022–2023), the detection distributions were 33.5% EAEC, 50.2% EPEC, and 12.5% ETEC, whereas during 2021–2022 they were 24.9% EAEC, 64.4% EPEC, and 6.2% ETEC (Fig. 1). Over time, the proportion of EAEC showed a continuous increasing trend (P < 0.05), whereas the proportion of EPEC gradually decreased.
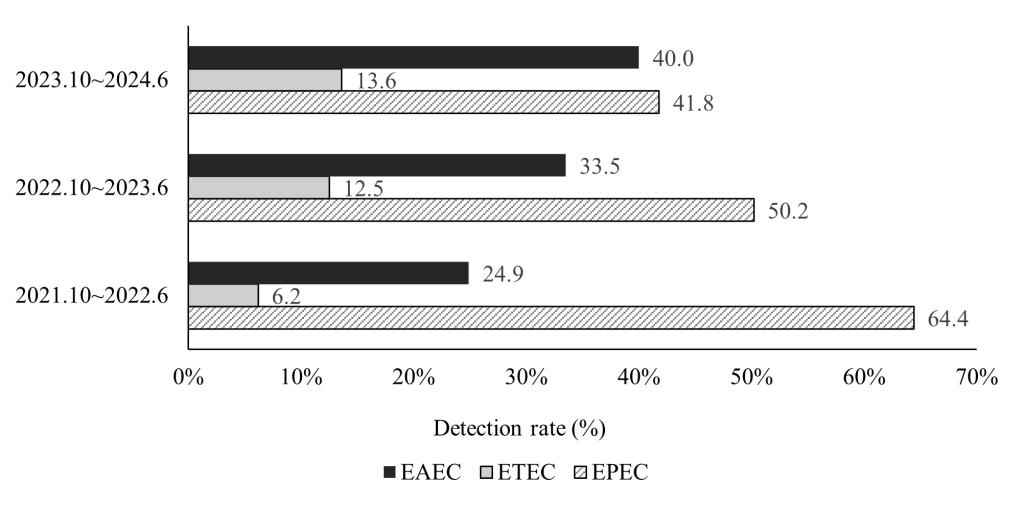
Fig. 1. Temporal trends in the prevalence of pathogenic Escherichia coli pathotypes (October 2021–June 2024). EAEC, enteroaggregative Escherichia coli; EPEC, enteropathogenic Escherichia coli; ETEC, enterotoxigenic Escherichia coli.
Our study identified EAEC as the most clinically significant pathotype among patients with diarrhea in the southeastern region of Korea, characterized by both high prevalence and concerning antibiotic resistance. Notably, EAEC exhibited the highest MDR rate (34.9%), distinguishing it as the predominant drug-resistant pathotype in the regional population. Furthermore, our findings revealed a significant age-dependent disparity, with pediatric patients showing higher resistance rates compared with adults.
The antibiotic resistance profiles of PEC isolates in this study showed high resistance to penicillins and cephalosporins, including AMP, CFZ, AMC, and CTX. These patterns are consistent with previous reports showing resistance rates exceeding 90% to penicillins and extensive resistance to β-lactams associated with the widespread clinical use of cephalosporins [4]. In addition to β-lactams, high resistance rates to SXT were observed, consistent with findings for both commensal and PEC in other regions [9,10]. Notably, CIP resistance was observed in 15.1% of all isolates, with an additional 49.5% showing intermediate susceptibility. This high prevalence of CIP non-susceptibility is concerning, given that E. coli is a major cause of urinary tract infections and fluoroquinolones are often used as a first-line treatment. The increased CIP resistance observed in our study, alongside high β-lactam resistance, poses a critical risk for treatment failure and recurrence, particularly in vulnerable populations [11]. Conversely, our study did not observe the high tetracycline resistance reported in some previous studies [5,6], likely reflecting the reduced clinical use of tetracycline and doxycycline in recent years. Recent studies have reported the spread of various tet gene variants in E. coli, leading to decreased tigecycline susceptibility [12]; however, resistance patterns remain variable by region.
When resistance rates were compared by pathotype, significant differences were observed among EAEC, EPEC, and ETEC. EAEC exhibited the highest resistance rates to AMP, AMC, CFZ, CTX, CAZ, and SXT. This finding suggests that EAEC has been exposed to stronger antibiotic selection pressures or possesses genetic characteristics that favor the accumulation of resistance genes [13,14]. EAEC is known to readily acquire resistance genes due to its diverse genetic composition and pathogenic plasmids that facilitate adherence to intestinal epithelial cells. Furthermore, biofilm formation by EAEC can prolong its persistence in the gut, increasing exposure to antibiotics and facilitating horizontal transfer of resistance genes [15]. In our study, EAEC displayed the most diverse resistance profiles, with concurrent resistance to β-lactams and SXT (e.g., AMP/SXT or AMP/CFZ/CTX combinations) occurring more frequently than in EPEC or ETEC. These characteristics suggest that EAEC should be prioritized in antibiotic management and surveillance systems, as healthy individuals carrying MDR-EAEC may serve as reservoirs for the spread of drug-resistant bacteria [16,17].
Comparison of antibiotic resistance rates by age group revealed that isolates from children had significantly higher resistance rates to β-lactams (AMP, AMC, CFZ, CTX, CAZ, and PT) than those from adults. In contrast, no significant differences were observed in SXT or CIP resistance between the two groups. Although reports directly comparing resistance rates by age are limited, our findings demonstrate that the antibiotic resistance burden may be higher among pediatric patients in this region. In addition, resistance rates to individual antibiotics and MDR prevalence were significantly higher in pediatric isolates than in adult isolates. These findings suggest a higher risk of MDR bacterial selection in the pediatric population, potentially related to frequent antibiotic use for common childhood infections.
This study has several limitations. First, because samples were collected solely from the Busan and Gyeongsangnam-do regions, the findings may reflect local antibiotic prescription patterns and PEC distribution, limiting direct generalization to nationwide trends. Second, the relatively small sample sizes for individual pathotypes limited the statistical power for detailed comparisons. Third, although we identified EAEC as a major resistant pathogen, further molecular studies are needed to elucidate the specific mechanisms of resistance and the genetic relatedness among these isolates.
Despite these limitations, our study provides valuable baseline data on the current state of antibiotic resistance among PEC in this region. The high prevalence of MDR-EAEC and the substantial resistance burden in pediatric patients underscore the need for continuous surveillance and the implementation of targeted antibiotic stewardship strategies.
This study was approved by the Institutional Review Board of Seegene medical foundation (IRB No. SMF-IRB-2023-021). Residual specimens from routine diagnostic testing were used, and the requirement for informed consent was waived by the IRB due to the retrospective nature of the study.
No potential conflicts of interest relevant to this article were reported.
None.
The datasets generated during the current study are available from the corresponding author upon request.
1. WHO. Diarrhoeal disease. https://www.who.int/news-room/fact-sheets/detail/diarrhoeal-disease [Online] (last visited on 7 March 2024).
2. GBD 2021 Diarrhoeal Diseases Collaborators. Global, regional, and national age-sex-specific burden of diarrhoeal diseases, their risk factors, and aetiologies, 1990-2021, for 204 countries and territories: a systematic analysis for the Global Burden of Disease Study 2021. Lancet Infect Dis 2025;25:519-36.![]()
![]()

3. Korea Disease Control and Prevention Agency. Guidelines for the management of water- and foodborne infectious diseases, 2026. https://www.kdca.go.kr/kdca/2861/subview.do?enc=Zm5jdDF8QEB8JTJGYmJzJTJGa2RjYSUyRjU1JTJGMzA5NTcxJTJGYXJ0Y2xWaWV3LmRvJTNG [Online] (last visited on 5 March 2026).
4. Kim YA, Park YS, Youk T, Lee H, Lee K. Trends in South Korean antimicrobial use and association with changes in Escherichia coli resistance rates: 12-year ecological study using a nationwide surveillance and antimicrobial prescription database. PLoS One 2018;13:e0209580.


5. Cho SH, Lim YS, Park MS, Kim SH, Kang YH. Prevalence of antibiotic resistance in Escherichia coli fecal isolates from healthy persons and patients with diarrhea. Osong Public Health Res Perspect 2011;2:41-5.


6. Park SB, Park YK, Ha MW, Thompson KD, Jung TS. Antimicrobial resistance, pathogenic, and molecular characterization of Escherichia coli from diarrheal patients in South Korea. Pathogens 2022;11:385.


7. CLSI. M100: Performance standards for antimicrobial susceptibility testing. 34th ed. Wayne, PA: Clinical and Laboratory Standards Institute; 2024.
8. Gupta K, Hooton TM, Naber KG, Wullt B, Colgan R, Miller LG, et al. International clinical practice guidelines for the treatment of acute uncomplicated cystitis and pyelonephritis in women: a 2010 update by the Infectious Diseases Society of America and the European Society for Microbiology and Infectious Diseases. Clin Infect Dis 2011;52:e103-20.

9. Nji E, Kazibwe J, Hambridge T, Joko CA, Larbi AA, Damptey LAO, et al. High prevalence of antibiotic resistance in commensal Escherichia coli from healthy human sources in community settings. Sci Rep 2021;11:3372.


10. Biswas S, Bal M, Pati S, Rana R, Dixit S, Ranjit M. Antibiotic resistance in toxigenic E. coli: a severe threat to global health. Discov Med 2024;1:72.
11. Flores-Mireles AL, Walker JN, Caparon M, Hultgren SJ. Urinary tract infections: epidemiology, mechanisms of infection and treatment options. Nat Rev Microbiol 2015;13:269-84.


12. Zhang S, Cui M, Liu D, Fu B, Shi T, Wang Y, et al. Tigecycline sensitivity reduction in Escherichia coli due to widely distributed tet(A) variants. Microorganisms 2023;11:3000.


13. Nüesch-Inderbinen MT, Hofer E, Hächler H, Beutin L, Stephan R. Characteristics of enteroaggregative Escherichia coli isolated from healthy carriers and from patients with diarrhea. J Med Microbiol 2013;62:1828-34.

14. Abdelwahab R, Yasir M, Godfrey RE, Christie GS, Element SJ, Saville F, et al. Antimicrobial resistance and gene regulation in Enteroaggregative Escherichia coli from Egyptian children with diarrhoea: similarities and differences. Virulence 2021;12:57-74.


15. Okeke IN and Nataro JP. Enteroaggregative Escherichia coli. Lancet Infect Dis 2001;1:304-13.

16. Tawfick MM, Elshamy AA, Mohamed KT, Menofy NGE. Gut commensal Escherichia coli, a high-risk reservoir of transferable plasmid-mediated antimicrobial resistance traits. Infect Drug Resist 2022;15:1077-91.


17. Purohit MR, Lindahl LF, Diwan V, Marrone G, Lundborg CS. High levels of drug resistance in commensal E. coli in a cohort of children from rural central India. Sci Rep 2019;9:6682.


1. WHO. Diarrhoeal disease. https://www.who.int/news-room/fact-sheets/detail/diarrhoeal-disease [Online] (last visited on 7 March 2024).
2. GBD 2021 Diarrhoeal Diseases Collaborators. Global, regional, and national age-sex-specific burden of diarrhoeal diseases, their risk factors, and aetiologies, 1990-2021, for 204 countries and territories: a systematic analysis for the Global Burden of Disease Study 2021. Lancet Infect Dis 2025;25:519-36.![]()
![]()

3. Korea Disease Control and Prevention Agency. Guidelines for the management of water- and foodborne infectious diseases, 2026. https://www.kdca.go.kr/kdca/2861/subview.do?enc=Zm5jdDF8QEB8JTJGYmJzJTJGa2RjYSUyRjU1JTJGMzA5NTcxJTJGYXJ0Y2xWaWV3LmRvJTNG [Online] (last visited on 5 March 2026).
4. Kim YA, Park YS, Youk T, Lee H, Lee K. Trends in South Korean antimicrobial use and association with changes in Escherichia coli resistance rates: 12-year ecological study using a nationwide surveillance and antimicrobial prescription database. PLoS One 2018;13:e0209580.


5. Cho SH, Lim YS, Park MS, Kim SH, Kang YH. Prevalence of antibiotic resistance in Escherichia coli fecal isolates from healthy persons and patients with diarrhea. Osong Public Health Res Perspect 2011;2:41-5.


6. Park SB, Park YK, Ha MW, Thompson KD, Jung TS. Antimicrobial resistance, pathogenic, and molecular characterization of Escherichia coli from diarrheal patients in South Korea. Pathogens 2022;11:385.


7. CLSI. M100: Performance standards for antimicrobial susceptibility testing. 34th ed. Wayne, PA: Clinical and Laboratory Standards Institute; 2024.
8. Gupta K, Hooton TM, Naber KG, Wullt B, Colgan R, Miller LG, et al. International clinical practice guidelines for the treatment of acute uncomplicated cystitis and pyelonephritis in women: a 2010 update by the Infectious Diseases Society of America and the European Society for Microbiology and Infectious Diseases. Clin Infect Dis 2011;52:e103-20.

9. Nji E, Kazibwe J, Hambridge T, Joko CA, Larbi AA, Damptey LAO, et al. High prevalence of antibiotic resistance in commensal Escherichia coli from healthy human sources in community settings. Sci Rep 2021;11:3372.


10. Biswas S, Bal M, Pati S, Rana R, Dixit S, Ranjit M. Antibiotic resistance in toxigenic E. coli: a severe threat to global health. Discov Med 2024;1:72.
11. Flores-Mireles AL, Walker JN, Caparon M, Hultgren SJ. Urinary tract infections: epidemiology, mechanisms of infection and treatment options. Nat Rev Microbiol 2015;13:269-84.


12. Zhang S, Cui M, Liu D, Fu B, Shi T, Wang Y, et al. Tigecycline sensitivity reduction in Escherichia coli due to widely distributed tet(A) variants. Microorganisms 2023;11:3000.


13. Nüesch-Inderbinen MT, Hofer E, Hächler H, Beutin L, Stephan R. Characteristics of enteroaggregative Escherichia coli isolated from healthy carriers and from patients with diarrhea. J Med Microbiol 2013;62:1828-34.

14. Abdelwahab R, Yasir M, Godfrey RE, Christie GS, Element SJ, Saville F, et al. Antimicrobial resistance and gene regulation in Enteroaggregative Escherichia coli from Egyptian children with diarrhoea: similarities and differences. Virulence 2021;12:57-74.


15. Okeke IN and Nataro JP. Enteroaggregative Escherichia coli. Lancet Infect Dis 2001;1:304-13.

16. Tawfick MM, Elshamy AA, Mohamed KT, Menofy NGE. Gut commensal Escherichia coli, a high-risk reservoir of transferable plasmid-mediated antimicrobial resistance traits. Infect Drug Resist 2022;15:1077-91.


17. Purohit MR, Lindahl LF, Diwan V, Marrone G, Lundborg CS. High levels of drug resistance in commensal E. coli in a cohort of children from rural central India. Sci Rep 2019;9:6682.

