1Department of Laboratory Medicine, Hallym University College of Medicine, Hallym University Hangang Sacred Heart Hospital, Seoul, Korea
2Department of Laboratory Medicine, Hallym University College of Medicine, Hallym University Kangnam Sacred Heart Hospital, Seoul, Korea
3Department of Laboratory Medicine, Hallym University College of Medicine, Hallym University Dongtan Sacred Heart Hospital, Hwaseong, Korea
4Department of Laboratory Medicine, Hallym University College of Medicine, Hallym University Sacred Heart Hospital, Anyang, Korea
5Department of Laboratory Medicine, Hallym University College of Medicine, Kangdong Sacred Heart Hospital, Seoul, Korea
Correspondence to Jae-Seok Kim E-mail: jaeseokcp@gmail.com
Ann Clin Microbiol 2026;29(1):1. https://doi.org/10.5145/ACM.2026.29.1.1
Received on 30 September 2025, Revised on 10 December 2025, Accepted on 27 January 2026, Published on 6 March 2026.
Copyright © Korean Society of Clinical Microbiology.
This is an Open Access article which is freely available under the Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License (CC BY-NC-ND) (https://creativecommons.org/licenses/by-nc-nd/4.0/).
Background: Enzyme immunoassays (EIAs), which detect glutamate dehydrogenase (GDH) and toxin A/B, are widely used to screen for Clostridioides difficile infection (CDI); however, their sensitivity is lower than that of molecular assays. This study aimed to evaluate the performance of two EIAs, C. Diff Quik Chek Complete (QCC) and RIDASCREEN (RIDA), and investigate the cycle threshold (Ct) values from two real-time polymerase chain reaction (PCR) assays (Allplex GI–Bacteria(I) and Xpert C. difficile) in EIA-discordant samples.
Methods: A total of 180 clinical stool samples were tested using QCC, RIDA, and Allplex GI-Bacteria(I) PCR assays. The Xpert C. difficile assay was used to analyze discordant results.
Results: QCC and RIDA showed high sensitivities for GDH detection, 100.0% and 94.4%, respectively. QCC was significantly more sensitive than RIDA for toxin detection (51.4% vs. 28.6%, p = 0.007). In 25 EIA-discordant, Xpert positive samples, the Ct values of the toxin B gene ranged from 31.5 to 44.8 (mean, 38.1) for Allplex PCR and from 23.7 to 36.3 (mean, 30.4) for Xpert PCR. The Ct values of the two PCR assays were not significantly correlated (r = 0.201, p = 0.324).
Conclusion: QCC is a suitable initial immunological test for diagnosing CDI. The lack of correlation in the Ct values between the two real-time PCR assays suggests that assay-specific validation is necessary for cutoff level interpretation.
Clostridioides difficile, Diff Quik, Glutamate dehydrogenase, Immunoenzyme techniques, Real-time polymerase chain reaction
Clostridioides difficile is the primary causative agent of antibiotic-associated diarrhea in healthcare facilities [1]. Excessive use of broad-spectrum antibiotics can disrupt the balance of the gut microbiome, resulting in an overgrowth of C. difficile leading to C. difficile infection (CDI) [2].
The diagnosis of CDI has evolved from traditional culture-based methods [3,4] to multi-step algorithms, which are now recommended by major guidelines such as the Infectious Diseases Society of America and the European Society of Clinical Microbiology and Infectious Diseases [5,6]. In Korea, a two-step algorithm is commonly employed, involving an initial screen using a sensitive glutamate dehydrogenase (GDH) assay and a toxin enzyme immunoassay (EIA), followed by a nucleic acid amplification test (NAAT) for discordant results (e.g., GDH-positive, toxin-negative) [7].
The effectiveness of such algorithms depends significantly on the performance of the initial EIA tests. Several commercial EIA platforms are available, and their performance characteristics may vary, thereby affecting the diagnostic accuracy. Therefore, the evaluation and comparison of these frontline assays are important. Furthermore, the use of NAATs has introduced new diagnostic challenges. While NAATs are highly sensitive, they can detect asymptomatic carriers, creating a clinically ambiguous scenario of a “NAATpositive, toxin EIA-negative” result [8].
To address this diagnostic uncertainty, an approach utilizing the real-time polymerase chain reaction (PCR) cycle threshold (Ct) value as a semi-quantitative proxy for bacterial load has emerged. The hypothesis that lower Ct values (indicating higher bacterial loads) correlate with more severe disease has been supported by some studies; however, the clinical utility of this approach remains debated [9–11]. There is a growing interest in the role of quantitative NAATs, particularly in Europe [12]; however, the comparability of Ct values generated by different molecular diagnostic platforms remains a critical, underinvestigated issue.
This study had two primary objectives. The first was to evaluate the performance of two widely used EIA platforms, C. Diff Quik Chek Complete (QCC; TechLab) and RIDASCREEN C. dif f icile GDH and Toxin A/B (RIDA; R–Biopharm). The second was to examine the correlation and compatibility of Ct values generated by two different real-time PCR platforms, the Seegene Allplex GI-Bacteria (I) assay (Seegene) and the Xpert C. difficile assay (Cepheid), in EIA-discordant samples.
We analyzed 180 residual stool samples that had been submitted for CDI testing at the Kangdong Sacred Heart Hospital, Seoul, Korea, between June 2019 and December 2019. After collection, all stool samples were stored at –70°C for 1 day to 6 months. All 180 samples were then thawed and tested simultaneously using QCC, RIDA, and the Allplex GI-Bacteria(I) assay. Subsequently, the samples were refrozen, and those showing discordant results were retested using the Xpert C. difficile assay within 3 days. This retrospective study evaluated the diagnostic accuracy, adhering to the Standards for Reporting Diagnostic Accuracy Studies guidelines (https://www.equator-network.org/reporting-guidelines/stard/).
C. Diff QCC is an immunochromatographic assay that simultaneously tests for GDH and toxins in a single cartridge. Stool samples were tested using QCC according to the manufacturer’s instructions. Each tube containing diluted stool sample was closed and thoroughly mixed. A 500-μL mixture, comprising 25 μL of stool sample with diluent and conjugate, was transferred to the device’s sample well. The mixture was left to incubate for 15 min at a temperature of 20°C–25°C. After incubation, 300 μL of wash buffer was added to the reaction window, followed by 2 drops of substrate. The results were recorded after 10 min. If a color band corresponding to GDH and/or toxins was visible in the display window of the device, it was reported to be positive.
RIDASCREEN C. difficile GDH and Toxin A/B (RIDA) are enzyme–linked immunosorbent assays that detect GDH and toxins A/B separately in 96-well plates. Stool samples were tested using RIDA according to the manufacturer’s instructions. RIDA GDH and RIDA Toxin A/B tests were performed sequentially using separate reagents. The GDH and toxin A/B concentrations were determined using an automated immunoassay system, EVOLIS (Bio-Rad), at a dual wavelength of 450 nm and 620 nm.
The Allplex GI-Bacteria (I) assay is a multiplex real-time PCR assay that can detect tcdB (encoding C. difficile toxin B). It also simultaneously detects other pathogens that cause gastrointestinal infections.
Stool samples were analyzed using the Allplex GI-Bacteria (I) assay in combination with the Microlab Nimbus system (Hamilton), which automated DNA extraction and PCR. A stool volume of 150–200 μL was transferred into 1 mL of stool lysis buffer. The suspension was pulse-vortexed for 1 min and incubated for an additional 10 min at room temperature. Afterward, the sample was centrifuged for 2 min at 14,000 rpm to obtain 800 µL of supernatant for the automated nucleic acid extraction procedure. PCR was performed on a CFX96 real time PCR detection system (Bio-Rad) as follows: 50°C for 20 min and 95°C for 15 min, followed by 45 cycles of 95°C for 10 s, 60°C for 10 s, and 72°C for 30 s. PCR results were interpreted using Seegene Viewer software (Seegene), and a positive result was defined as a fluorescence curve crossing the Ct at a value of < 45.
The Xpert C. difficile assay is a fully automated, integrated, closed system that detects tcdB and binary toxin, as well as the deletion at nucleotide 117 on tcdC, the gene encoding the negative regulator of toxin production, which serves as a marker for identifying a C. difficile virulent strain known as 027/NAP1/BI [13].
Stool samples were tested using the Xpert C. difficile assay according to the manufacturer’s instructions. Briefly, a swab was taken from each stool sample and inserted into a vial containing sample reagent. The vials were then vortexed at high speed for 10 s. The entire sample was transferred to the sample chamber of the Xpert C. difficile assay cartridge. The results were interpreted using the GeneXpert Dx system (Cepheid), and a positive result was defined as a Ct value of < 38.
Samples positive in both molecular assays were considered true positives, and those negative in both assays were considered true negatives. In cases of discordant results, the final classification was determined according to the Xpert C. difficile assay result. The sensitivity, specificity, positive predictive value (PPV), negative predictive value (NPV), positive likelihood ratio, negative likelihood ratio, area under the curve (AUC), and 95% confidence interval (CI) were calculated. CIs for sensitivity, specificity, and accuracy were calculated using the Clopper–Pearson exact method. CIs for the likelihood ratios were calculated using the “Log method” [14]. The CIs for predictive values are standard logit CIs [15], except when the predictive value is 0 or 100.0%, in which a Clopper–Pearson CI is reported [16]. The DeLong test was used to compare the AUCs [17]. The correlation between the Ct values of the two molecular methods was analyzed using the Pearson correlation analysis. A p–value < 0.05 was considered statistically significant. Statistical analyses were performed using SPSS version 24 (IBM Corp.) and MedCalc version 10.0 (MedCalc Software Ltd.).
The concordance rates of QCC and RIDA-GDH for true positives and true negatives were 93.9% (169/180) and 89.4% (161/180), respectively. QCC/RIDA GDH demonstrated sensitivities of 100.0% and 94.4%, specificities of 92.4% and 88.2%, PPV of 76.6% and 66.7%, and NPV of 100.0% and 98.4%, respectively (Fig. 1, Table 1). QCC and RIDA GDH showed comparable AUCs (QCC, 0.959; 95% CI, 0.918–0.983 vs. RIDA GDH, 0.927; 95% CI, 0.879–0.960; p = 0.073; Fig. 2).
The concordance rates of QCC and RIDA toxins for true positives and true negatives were 90.0% (162/180) and 86.1% (155/180), respectively. The QCC/RIDA toxin demonstrated sensitivities of 51.4% and 28.6%, specificities of 99.3% and 100.0%, PPV of 94.7% and 100.0%, and NPV of 89.4% and 85.3%, respectively (Fig. 1, Table 1). QCC showed significantly higher AUC than that of RIDA toxin (QCC, 0.736; 95% CI, 0.665–0.799 vs. RIDA toxin, 0.643; 95% CI, 0.568–0.713; p = 0.007; Fig. 2).
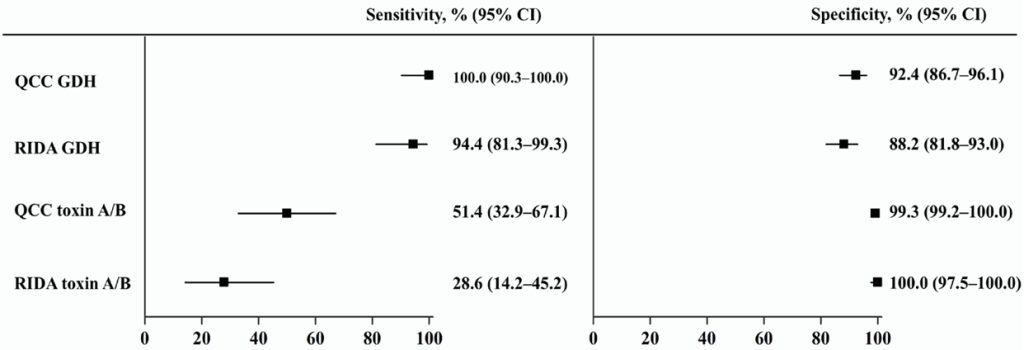
Fig. 1. Sensitivity and specificity of C. Diff Quik Chek Complete and RIDASCREEN C. difficile GDH and toxin A/B. QCC, C. Diff Quik Chek Complete; RIDA, RIDASCREEN C. difficile assay; X-axis, percentages.
Table 1. The performance of C. Diff Quik Chek Complete and RIDASCREEN C. difficile GDH and toxin A/B
| Sensitivity (95% CI) | Specificity (95% CI) | Positive likelihood ratio (95% CI) | Negative likelihood ratio (95% CI) | AUC (95% CI, P) | PPV (95% CI) | NPV (95% CI) | Accuracy (95% CI) | |
|---|---|---|---|---|---|---|---|---|
| QCC GDH | 100.0% (90.3–100.0) | 92.4% (86.7–96.1) | 13.1 (7.4–23.1) | 0.0 | 0.959 (0.918–0.983, <0.001) | 76.6% (65.0–85.2) | 100.0% (97.3–100.0) | 93.9% (89.3–96.9) |
| RIDA GDH | 94.4% (81.3–99.3) | 88.2% (81.8–93.0) | 8.0 (5.1–12.6) | 0.1 (0.0–0.2) | 0.927 (0.879–0.960, <0.001) | 66.7% (56.0–75.9) | 98.4% (94.3–99.6) | 89.4% (84.0–93.5) |
| QCC toxin A/B | 51.4% (34.0–68.6) | 99.3% (96.2–100.0) | 74.6 (10.3–539.9) | 0.5 (0.4–0.7) | 0.736 (0.665–0.799, <0.001) | 94.7% (71.3–99.2) | 89.4% (85.8–92.2) | 90.0% (84.7–94.0) |
| RIDA toxin A/B | 28.6% (14.6–46.3) | 100.0% (97.5–100.0) | – | 0.7 (0.6–0.9) | 0.643 (0.568–0.713, <0.001) | 100.0% (69.2–100.0) | 85.3% (82.5–87.7) | 86.1% (80.2–90.8) |
Abbreviations: GDH, glutamate dehydrogenase; CI, confidence interval; AUC, area under the curve; PPV, positive predictive value; NPV, negative predictive value; QCC, C. Diff Quik Chek Complete; RIDA, RIDASCREEN C. difficile assay.
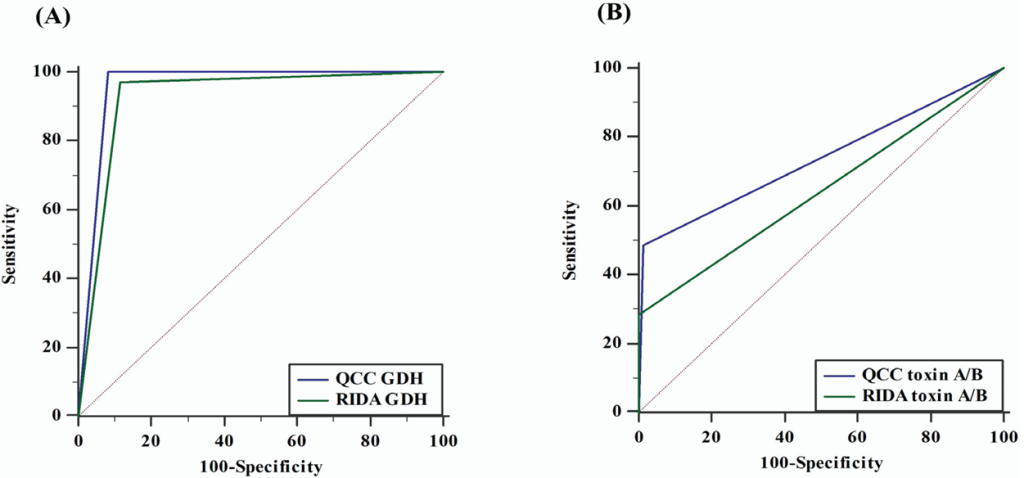
Fig. 2. Area under curve (AUC) of C. Diff Quik Chek Complete and RIDASCREEN C. difficile GDH and toxin A/B. (A) AUC of GDH; (B) AUC of toxin A/B. QCC, C. Diff Quik Chek Complete; RIDA, RIDASCREEN C. difficile assay; GDH, glutamate dehydrogenase.
Of the 180 stool samples collected, 36 (20.0%) tested positive in the Allplex GI-Bacteria(I) assay. Among the 36 positive samples, 26 showed discordant QCC and RIDA toxin results. These 26 samples were tested using the Xpert C. difficile assay, 25 of which showed positive results (Table 2).
The Ct values of the 25 “true-positive” samples ranged from 31.5 to 44.8 in the Allplex GI-Bacteria(I) assay and from 23.7 to 36.3 in the Xpert C. difficile assay (Fig. 3). The Ct value of one sample that tested negative in the Xpert C. difficile assay was 35.5 in the Allplex GI-Bacteria(I) assay and 38.1 in the Xpert C. difficile assay. No significant correlation was observed between Ct values of the two molecular assays (r = 0.201, p = 0.324).
Table 2. Results of Allplex GI-Bacteria(I) and Xpert C. difficile assays in 26 EIA-discordant samples
| QCC | RIDA | Allplex GI-Bacteria(I) assay (Toxin B) | Xpert C. difficile assay (Toxin B) | No. of specimens | ||
|---|---|---|---|---|---|---|
| GDH | Toxin A/B | GDH | Toxin A/B | |||
| Positive | Negative | Positive | Negative | Positive | Positive | 16 |
| Positive | Positive | Positive | Negative | Positive | Positive | 7 |
| Positive | Negative | Negative | Negative | Positive | Positive | 1 |
| Positive | Positive | Negative | Negative | Positive | Positive | 1 |
| Positive | Negative | Positive | Negative | Positive (Ct 35.5) | Negative (Ct 38.1)a) | 1 |
a)The cutoff Ct value for toxin B in the Xpert C. difficile assay was 38.0. However, the Ct value near the cutoff value of the toxin B gene can be used for further evaluation.
Abbreviations: QCC, C. Diff Quik Chek Complete; RIDA, RIDASCREEN C. difficile assay; GDH, glutamate dehydrogenase; Ct, cycle threshold.
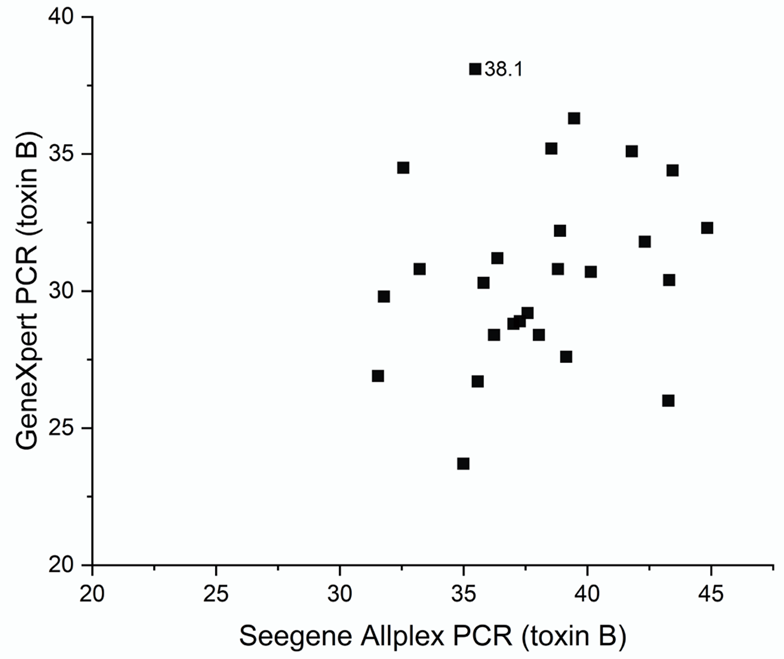
Fig. 3. Correlation of toxin B gene cycle threshold values from Allplex GI-Bacteria(I) and Xpert C. difficile assays. PCR, polymerase chain reaction.
QCC and RIDA GDH demonstrated high concordance (93.9%, 89.4%) with sensitivities of 100.0% and 94.4%, respectively, and comparable AUC. For toxin detection, QCC showed significantly higher sensitivity than RIDA (51.4% vs 28.6%) and superior AUC (0.736 vs 0.643, p=0.007). Among 180 samples, 36 (20%) were Allplex-positive. Of 26 EIA toxin-discordant samples, 25 were Xpert-positive. True-positive Ct values ranged from 31.5–44.8 (Allplex) and 23.7–36.3 (Xpert).
In this study, both QCC and RIDA showed high sensitivity and specificity for GDH detection, consistent with previous reports [18–21]. However, a significant performance difference was observed for toxin A/ B detection. The sensitivity of QCC (51.4%) was consistent with previous studies [6,20–22], whereas the sensitivity of RIDA was notably lower at 28.6%, which fell below the previously reported range of 33.0%86.0% [7,20,21,23]. This suggests that RIDA may not have detected a substantial proportion of true CDI cases in this study population.
Although we acknowledge that the use of stored stool samples can affect toxin stability [24], this finding does not appear to be an isolated phenomenon. In a similar study by Yoo et al., which used toxigenic culture as the reference method, the RIDA toxin assay also showed lower sensitivity (33.3%) than the QCC assay (60.0%) [20]. This suggests that the difference in performance between the assays is a consistent finding beyond the limitations of our sample conditions. Therefore, based on its superior toxin detection sensitivity, QCC is a more suitable choice for the initial screening test in a multistep CDI diagnostic algorithm.
The use of real-time PCR Ct values as a proxy for the bacterial load has shown significant promise for improving CDI diagnostics. Kamboj et al. [11] reported that a Ct value cutoff of 28.0 on the Xpert assay could predict 91.0% of severe and 100.0% of complicated CDI cases, establishing a direct link between lower Ct values and increased clinical severity. Furthermore, Suleiman et al. [8] demonstrated that incorporating Ct value reporting into the diagnostic process reduced unnecessary antibiotic treatment by 23% (p = 0.025) in pediatric populations. Although these studies identified specific Ct value cutoffs using the Xpert assay, other commercial platforms, such as the Allplex system, are also used in clinical settings. Consequently, there is a risk that clinicians or laboratories may attempt to apply published Ct cutoffs derived from Xpert studies to results obtained from other platforms. However, the results of our study revealed a fundamental problem with this Ct-value-based approach for risk stratification.
The core finding of our study was the lack of statistically significant correlation (r = 0.201, p = 0.324) between the Ct values obtained from the Seegene Allplex and Xpert assays on the same set of discordant samples. This discrepancy can be attributed to various technical factors, including differences in DNA extraction protocols, primer and probe sequences targeting different gene regions, amplification efficiencies, and proprietary algorithms used by each system to determine the Ct. Thus, the Ct value of a sample obtained from one platform cannot be reliably extrapolated to an equivalent value on the same sample obtained from another platform. This finding challenges the feasibility of establishing a universal Ct cutoff for clinical decision-making because applying a cutoff value validated for one assay to the results from another assay could lead to inconsistent risk stratification. Therefore, our results underscore the necessity of platform-specific validation before generalized Ct value cutoffs are broadly adopted in clinical guidelines.
The clinical management of patients with NAAT-positive/toxin-negative status remains debatable. While these patients may have a lower risk of recurrence [25,26], recent meta-analyses show that their 30-day mortality is often not statistically different from patients with NAAT-positive/toxin-positive [25,26]. This suggests that a subset of this group remains at risk for severe outcomes, highlighting the potential danger of withholding treatment based solely on negative toxin EIA results.
Our results indicate the lack of interchangeability in Ct values between PCR platforms, especially in samples with high Ct values, and a universal and reliable Ct value cutoff to stratify risk in this population may not be feasible. This finding reinforces the value of the proposed multistep algorithm. Because a universal Ct cutoff value cannot be reliably applied, using a sensitive NAAT to clarify discordant initial EIA results is essential to provide clinicians with the critical information needed for these nuanced decisions.
This study had several limitations. First, although toxigenic culture or cell cytotoxicity neutralization assay has traditionally been regarded as the reference standard, many recent studies have used NAAT-based methods as practical reference standards. Nevertheless, the absence of toxigenic culture may still have influenced the estimated performance of EIAs. Second, our sample collection was limited to a single 500-bed hospital, and the relatively small sample size of 180 stool samples may have affected the results. Third, with regard to specimen stability, the Xpert C. difficile assay was not conducted simultaneously with the initial EIAs or Allplex assay. Although samples were stored at –70°C to minimize degradation, the interval between tests and the freeze-thaw process could potentially impact the detection of labile toxins or nucleic acid integrity. Although DNA is generally more stable than protein toxins, the lack of correlation in Ct values between Seegene Allplex and Xpert assays could be influenced by the storage duration; however, the consistent internal controls in both molecular assays suggested that degradation was not the primary driver of the observed discrepancies. Nevertheless, we attempted to minimize this limitation by ensuring standardized storage conditions. Fourth, the absence of clinical information from the patients precluded evaluation of the test results for clinical plausibility.
QCC demonstrated superior toxin detection sensitivity compared to RIDA, making it more suitable for the initial screening step of a CDI diagnostic algorithm. The Ct values for the C. difficile toxin B gene were not interchangeable between the Seegene Allplex and Xpert assays. Our results underscore the need for platform-specific validation. A universal Ct cutoff for CDI risk stratification is currently unachievable, highlighting a critical area for future research and standardization before such strategies can be safely and effectively implemented in clinical practice.
This study was approved by the Institutional Review Board (IRB) of Kangdong Sacred Heart Hospital (IRB No. 2019–07–016), which waived the requirement for informed consent.
No potential conflicts of interest relevant to this article were reported.
This research was supported by the Ministry of Trade, Industry and Energy (Technology Innovation Program, Grant No. 10047748).
The datasets generated during the current study are available from the corresponding author upon request.
1. Evans CT and Safdar N. Current trends in the epidemiology and outcomes of Clostridium difficile infection. Clin Infect Dis 2015;60 Suppl 2:S66-71.

2. Taur Y and Pamer EG. Harnessing microbiota to kill a pathogen: fixing the microbiota to treat Clostridium difficile infections. Nat Med 2014;20:246-7.


3. Planche TD, Davies KA, Coen PG, Finney JM, Monahan IM, Morris KA, et al. Differences in outcome according to Clostridium difficile testing method: a prospective multicentre diagnostic validation study of C difficile infection. Lancet Infect Dis 2013;13:936-45.


4. Azrad M, Tkhawkho L, Hamo Z, Peretz A. The diagnostic performance and accuracy of 3 molecular assays for the detection of Clostridium difficile in stool samples, compared with the Xpert® C. difficile assay. J Microbiol Methods 2020;168:105784.

5. McDonald L, Gerding D, Johnson S, Bakken J, Carroll K, Coffin S, et al. Clinical practice guidelines for Clostridium difficile infection in adults and children: 2017 update by the Infectious Diseases Society of America (IDSA) and Society for Healthcare Epidemiology of America (SHEA). Clin Infect Dis 2018;66:987-94.


6. Crobach M, Planche T, Eckert C, Barbut F, Terveer E, Dekkers O, et al. European Society of Clinical Microbiology and Infectious Diseases: update of the diagnostic guidance document for Clostridium difficile infection. Clin Microbiol Infect 2016;22:S63-81.

7. Chung HS, Park JS, Shin BM, Yoo HM, Kim H, Cho J, et al. Nationwide survey for current status of laboratory diagnosis of Clostridioides difficile infection in Korea. J Korean Med Sci 2022;37:e38.


8. Suleiman M, Tang P, Imam O, Morales P, Altrmanini D, Barr KL, et al. Use of PCR cycle threshold and clinical interventions to aid in the management of pediatric Clostridioides difficile patients. Microorganisms 2024;12:1181.


9. Smith-Aguasca R, Camenforte N, Rodríguez L, Herrera S, Vara K, Lueerssen D, et al. Use of PCR cycle threshold values for toxins A and B quantification in Clostridioides difficile infections. Sci Rep 2025;15:18730.


10. Lee S, Nanda N, Yamaguchi K, Lee Y, She RC. Clostridioides difficile toxin B PCR cycle threshold as a predictor of toxin testing in stool specimens from hospitalized adults. Antibiotics 2022;11:576.


11. Kamboj M, Brite J, McMillen T, Robilotti E, Herrera A, Sepkowitz K, et al. Potential of real-time PCR threshold cycle (CT) to predict presence of free toxin and clinically relevant C. difficile infection (CDI) in patients with cancer. J Infect 2018;76:369-75.


12. European Centre for Disease Prevention and Control. Laboratory procedures for diagnosis and typing of human Clostridium difficile infection. Stockholm, Sweden: ECDC; 2018.
13. Somily AM, Khan MA, Morshed M. The laboratory diagnosis of Clostridioides difficile infection: an update of current laboratory practice. J Infect Dev Ctries 2021;15:1364-75.

14. Altman DG, Machin D, Bryant TN, Gardner MJ. Statistics with confidence. 2nd ed. London: BMJ Books; 2001:240.
15. Mercaldo ND, Lau KF, Zhou XH. Confidence intervals for predictive values with an emphasis to case-control studies. Stat Med 2007;26:2170-83.

16. MedCalc Software Ltd. MedCalc Statistical Software (version 18.9.1). https://www.medcalc.org [Online] (last visited on 7 May 2024).
17. DeLong ER, DeLong DM, Clarke-Pearson DL. Comparing the areas under two or more correlated receiver operating characteristic curves: a nonparametric approach. Biometrics 1988;44:837-45.

18. Ota KV and McGowan KL. Clostridium difficile testing algorithms using glutamate dehydrogenase antigen and C. difficile toxin enzyme immunoassays with C. difficile nucleic acid amplification testing increase diagnostic yield in a tertiary pediatric population. J Clin Microbiol 2012;50:1185-8.


19. Swindells J, Brenwald N, Reading N, Oppenheim B. Evaluation of diagnostic tests for Clostridium difficile infection. J Clin Microbiol 2010;48:606-8.


20. Yoo IY, Song DJ, Huh HJ, Lee NY. Simultaneous detection of Clostridioides difficile glutamate dehydrogenase and toxin A/B: comparison of the C. DIFF QUIK CHEK COMPLETE and RIDASCREEN assays. Ann Lab Med 2019;39:214-7.


21. Carroll KC and Mizusawa M. Laboratory tests for the diagnosis of Clostridium difficile. Clin Colon Rectal Surg 2020;33:73-81.


22. Chung HS and Lee M. Evaluation of the performance of C. DIFF QUIK CHEK COMPLETE and its usefulness in a hospital setting with a high prevalence of Clostridium difficile infection. J Investig Med 2017;65:88-92.

23. Mattner F, Winterfeld I, Mattner L. Diagnosing toxigenic Clostridium difficile: new confidence bounds show culturing increases sensitivity of the toxin A/B enzyme immunoassay and refute gold standards. Scand J Infect Dis 2012;44:578-85.

24. Vohra P and Poxton IR. Comparison of toxin and spore production in clinically relevant strains of Clostridium difficile. Microbiology 2011;157:1343-53.

25. Prosty C, Hanula R, Katergi K, Longtin Y, McDonald EG, Lee TC. Clinical outcomes and management of NAAT-positive/Toxin-negative Clostridioides difficile infection: a systematic review and meta-analysis. Clin Infect Dis 2024;78:430-8.

26. Tansarli GS, Falagas ME, Fang FC. Clinical significance of toxin EIA positivity in patients with suspected Clostridioides difficile infection: systematic review and meta-analysis. J Clin Microbiol 2025;63:e00977-24.


1. Evans CT and Safdar N. Current trends in the epidemiology and outcomes of Clostridium difficile infection. Clin Infect Dis 2015;60 Suppl 2:S66-71.

2. Taur Y and Pamer EG. Harnessing microbiota to kill a pathogen: fixing the microbiota to treat Clostridium difficile infections. Nat Med 2014;20:246-7.


3. Planche TD, Davies KA, Coen PG, Finney JM, Monahan IM, Morris KA, et al. Differences in outcome according to Clostridium difficile testing method: a prospective multicentre diagnostic validation study of C difficile infection. Lancet Infect Dis 2013;13:936-45.


4. Azrad M, Tkhawkho L, Hamo Z, Peretz A. The diagnostic performance and accuracy of 3 molecular assays for the detection of Clostridium difficile in stool samples, compared with the Xpert® C. difficile assay. J Microbiol Methods 2020;168:105784.

5. McDonald L, Gerding D, Johnson S, Bakken J, Carroll K, Coffin S, et al. Clinical practice guidelines for Clostridium difficile infection in adults and children: 2017 update by the Infectious Diseases Society of America (IDSA) and Society for Healthcare Epidemiology of America (SHEA). Clin Infect Dis 2018;66:987-94.


6. Crobach M, Planche T, Eckert C, Barbut F, Terveer E, Dekkers O, et al. European Society of Clinical Microbiology and Infectious Diseases: update of the diagnostic guidance document for Clostridium difficile infection. Clin Microbiol Infect 2016;22:S63-81.

7. Chung HS, Park JS, Shin BM, Yoo HM, Kim H, Cho J, et al. Nationwide survey for current status of laboratory diagnosis of Clostridioides difficile infection in Korea. J Korean Med Sci 2022;37:e38.


8. Suleiman M, Tang P, Imam O, Morales P, Altrmanini D, Barr KL, et al. Use of PCR cycle threshold and clinical interventions to aid in the management of pediatric Clostridioides difficile patients. Microorganisms 2024;12:1181.


9. Smith-Aguasca R, Camenforte N, Rodríguez L, Herrera S, Vara K, Lueerssen D, et al. Use of PCR cycle threshold values for toxins A and B quantification in Clostridioides difficile infections. Sci Rep 2025;15:18730.


10. Lee S, Nanda N, Yamaguchi K, Lee Y, She RC. Clostridioides difficile toxin B PCR cycle threshold as a predictor of toxin testing in stool specimens from hospitalized adults. Antibiotics 2022;11:576.


11. Kamboj M, Brite J, McMillen T, Robilotti E, Herrera A, Sepkowitz K, et al. Potential of real-time PCR threshold cycle (CT) to predict presence of free toxin and clinically relevant C. difficile infection (CDI) in patients with cancer. J Infect 2018;76:369-75.


12. European Centre for Disease Prevention and Control. Laboratory procedures for diagnosis and typing of human Clostridium difficile infection. Stockholm, Sweden: ECDC; 2018.
13. Somily AM, Khan MA, Morshed M. The laboratory diagnosis of Clostridioides difficile infection: an update of current laboratory practice. J Infect Dev Ctries 2021;15:1364-75.

14. Altman DG, Machin D, Bryant TN, Gardner MJ. Statistics with confidence. 2nd ed. London: BMJ Books; 2001:240.
15. Mercaldo ND, Lau KF, Zhou XH. Confidence intervals for predictive values with an emphasis to case-control studies. Stat Med 2007;26:2170-83.

16. MedCalc Software Ltd. MedCalc Statistical Software (version 18.9.1). https://www.medcalc.org [Online] (last visited on 7 May 2024).
17. DeLong ER, DeLong DM, Clarke-Pearson DL. Comparing the areas under two or more correlated receiver operating characteristic curves: a nonparametric approach. Biometrics 1988;44:837-45.

18. Ota KV and McGowan KL. Clostridium difficile testing algorithms using glutamate dehydrogenase antigen and C. difficile toxin enzyme immunoassays with C. difficile nucleic acid amplification testing increase diagnostic yield in a tertiary pediatric population. J Clin Microbiol 2012;50:1185-8.


19. Swindells J, Brenwald N, Reading N, Oppenheim B. Evaluation of diagnostic tests for Clostridium difficile infection. J Clin Microbiol 2010;48:606-8.


20. Yoo IY, Song DJ, Huh HJ, Lee NY. Simultaneous detection of Clostridioides difficile glutamate dehydrogenase and toxin A/B: comparison of the C. DIFF QUIK CHEK COMPLETE and RIDASCREEN assays. Ann Lab Med 2019;39:214-7.


21. Carroll KC and Mizusawa M. Laboratory tests for the diagnosis of Clostridium difficile. Clin Colon Rectal Surg 2020;33:73-81.


22. Chung HS and Lee M. Evaluation of the performance of C. DIFF QUIK CHEK COMPLETE and its usefulness in a hospital setting with a high prevalence of Clostridium difficile infection. J Investig Med 2017;65:88-92.

23. Mattner F, Winterfeld I, Mattner L. Diagnosing toxigenic Clostridium difficile: new confidence bounds show culturing increases sensitivity of the toxin A/B enzyme immunoassay and refute gold standards. Scand J Infect Dis 2012;44:578-85.

24. Vohra P and Poxton IR. Comparison of toxin and spore production in clinically relevant strains of Clostridium difficile. Microbiology 2011;157:1343-53.

25. Prosty C, Hanula R, Katergi K, Longtin Y, McDonald EG, Lee TC. Clinical outcomes and management of NAAT-positive/Toxin-negative Clostridioides difficile infection: a systematic review and meta-analysis. Clin Infect Dis 2024;78:430-8.

26. Tansarli GS, Falagas ME, Fang FC. Clinical significance of toxin EIA positivity in patients with suspected Clostridioides difficile infection: systematic review and meta-analysis. J Clin Microbiol 2025;63:e00977-24.

