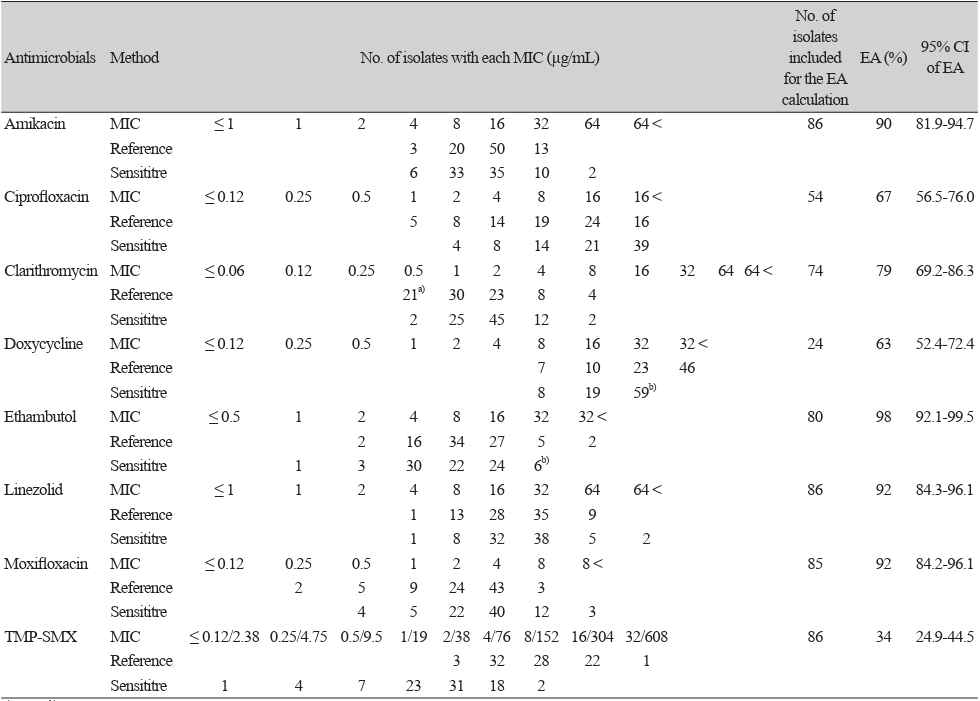
Drug susceptibility testing of Mycobacterium avium complex using the SLOMYCO test-system: a diagnostic accuracy study
Original article Jeong Su Park1*, Kyu-Hwa Hur2*, Woo Jin Shin1, Hyunji Kim1, Dong Woo Shin1, Kyoung Un Park1 1Department of Laboratory Medicine, Seoul National University Bundang Hospital, Seoul National University College of Medicine, Seongnam, Korea2Department of Laboratory Medicine, Green Cross Laboratories, Yongin, Korea*These authors contributed equally to this work. Correspondence to Jeong Su Park, E-mail: […]
Prevalence and molecular characteristics of β-lactam resistance in non-typeable Haemophilus influenzae isolates in Korea
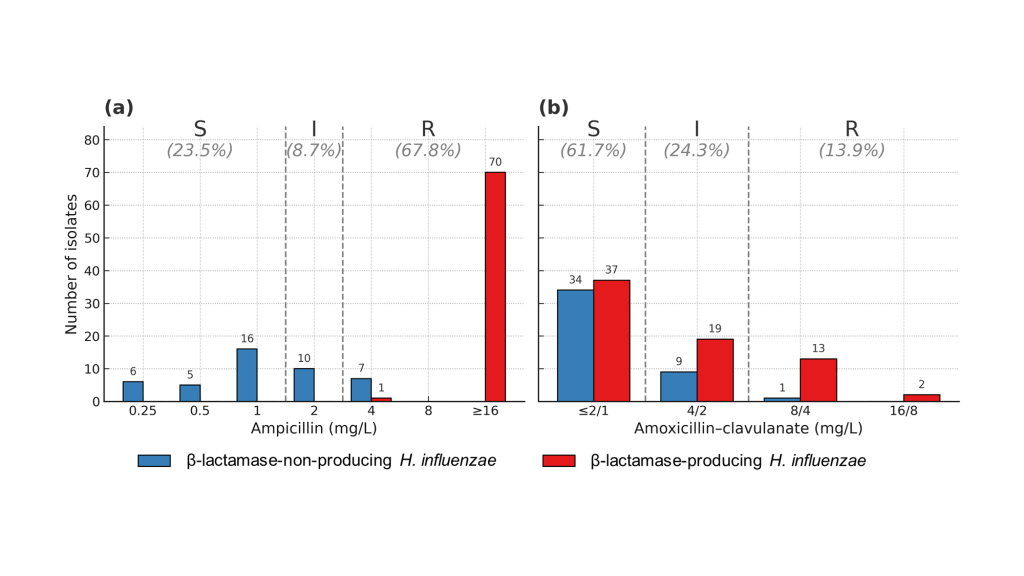
Original article Eun-Young Kim1, Yeon Chan Choi1, Hyeon Jin Choi1, Si Hyun Kim2, Jihyun Cho3, Seok Hoon Jeong4, Dokyun Kim4, Hyun Soo Kim5, Soo Hyun Kim6, Young Ah Kim7, Young Ree Kim8, Nam Hee Ryoo9, Jong Hee Shin6, Kyeong Seob Shin10, Young Uh11, Jeong Hwan Shin1 1Department of Laboratory Medicine and Paik Institute for Clinical […]
Fig. 4. Putative identification of other streptococcal ICEs resembling Streptococcus uberis ICEs (UB37/UB68/UB23/UB99) with the S. uberis NZ01 reference genome. Similar or identical products between reference and candidate genomes are indicated by matching colors. Black gradations between corresponding genes indicate percent identity. A similar streptococcal ICE was identified in Streptococcus suis strain STC78 (ICEnsui78–tet(O)–erm(B); accession number ON944185), based on conserved product arrangements between UB37/UB68/UB23/UB99 ICEs and ICEnsui78. Core products are in red font, and antimicrobial resistance products are in blue font. ICEUB37 was re-annotated using the Prokaryotic Genomes Annotation Pipeline, and ICE components were identified using ICEscreen. Protein function similarities were inferred using Basic Local Alignment Search Tool x, with the predicted protein functions shown in brackets. Figure is cited from [45]. WGS, whole-genome sequencing; ICE, integrative and conjugative element.
Ann Clin Microbiol 2025;28(4):22. Whole-genome sequencing applications for evolution of clinical microbiology Download image
Fig. 3. Bacterial RNA isolation workflow. Mechanical lysis was performed using a MagNA lyser (Roche). Human keratinocytes were lysed with 1.4 mm silica beads (Qbiogene) in RLT lysis buffer (RNeasy Fibrous Tissue Mini Kit; Qiagen), and the human RNA fraction was removed by centrifugation. The pellets were then lysed with 0.1 mm silica beads (Qbiogene) in RLT lysis buffer and centrifuged to obtain the bacterial RNA fraction. Figure cited from [29].
Ann Clin Microbiol 2025;28(4):22. Whole-genome sequencing applications for evolution of clinical microbiology Download image
Fig. 2. Genome structure containing the cytolethal distending toxin (cdt)A–cdtB–cdtC loci of Pasteurella canis and adjacent loci from strain HL_NV12211 (accession number CP085871), compared to the corresponding region of Pasteurella multocida subsp. multocida ATCC 43137(T) (accession number CP008918). Asterisks indicate putative Holliday junction resolvase. K7G93_001965, K7G93_001967, and DR93_66 represent loci encoding hypothetical proteins. relA, GTP diphosphokinase; rlmD and rumA, 23S rRNA (uracil(1939)-C(5))-methyltransferase; recO, DNA repair protein; rsmE, 16S rRNA (uracil(1498)-N(3))-methyltransferase; eno, phosphopyruvate hydratase; pyrG, CTP synthase. Figure cited from [25].
Ann Clin Microbiol 2025;28(4):22. Whole-genome sequencing applications for evolution of clinical microbiology Download image
Fig. 1. Clinker visualization. Genes within a gene cluster are color-coded, and identical or similar genes among multiple clusters are connected by links shaded based on sequence identity. Figure cited from [24].
Ann Clin Microbiol 2025;28(4):22. Whole-genome sequencing applications for evolution of clinical microbiology Download image
Table 3. Commonly used antimicrobial resistance databases [40]
Ann Clin Microbiol 2025;28(4):22. Whole-genome sequencing applications for evolution of clinical microbiology Download table Database [curator] Traits AMRFinderPlus Database [NCBI] Comprehensive and curated database used by NCBI’s AMRFinderPlus software toolsResponsible for designation of allele names for new beta-lactamase and tetracycline-resistant genes Comprehensive Antimicrobial Resistance Database (CARD) [McMaster University] Comprehensive database of AMR gene sequences and […]
Table 2. Comparison of strengths and weaknesses of traditional approaches and whole-genome sequencing [3]
Ann Clin Microbiol 2025;28(4):22. Whole-genome sequencing applications for evolution of clinical microbiology Download table Item Traditional approaches Whole-genome sequencing Principle Phenotypic traits, such as culturing, serotyping, biochemical testing, or PCR-based detection Sequencing the entire genome to identify pathogens and analyze genetic features Applications Detection, identification, and enumeration of pathogens Outbreak tracing, source attribution, evolution study, […]
Table 1. Historical timeline of discoveries in molecular diagnostic procedures, with descriptions and applications in the 21st century [1]
Ann Clin Microbiol 2025;28(4):22. Whole-genome sequencing applications for evolution of clinical microbiology Download table Year Molecular procedures Description Applications 2000s Microarrays Analysis of gene expression, SNP genotyping, and comparative genomic hybridization Study of gene expression, detection of genetic variation, and identification of chromosomal abnormalities 2000s Metagenomics Comprehensive analysis of entire pathogen populations Identification of rare […]
Table 2.Cycle threshold values and melt peak temperatures of Xpert MTB/RIF and Xpert MTB/RIF Ultra assays
Ann Clin Microbiol 2025;28(4):21. Impact of nontuberculous mycobacteria on the performance of Xpert MTB/RIF and Xpert MTB/RIF Ultra for the detection of tuberculosis and rifampin resistance: a diagnostic accuracy study Download table Concentration(CFU/mL equivalent) Xpert MTB/RIF Xpert MTB/RIF Ultra Ct value (1st/2nd) Ct value (1st/2nd) Tm (1st/2nd) NTM strain(1.0 × 106) MTB(5.0×103) […]
